[English] 日本語
 Yorodumi
Yorodumi- EMDB-38245: Open State of central tail fiber of bacteriophage lambda upon bin... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Open State of central tail fiber of bacteriophage lambda upon binding to LamB (gpJ713-LamB complex) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacteriophage / caudovirales / siphoviridae / phage lambda / host recognition / LamB / cryo-EM / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationmaltodextrin transmembrane transporter activity / maltose transporting porin activity / symbiont genome ejection through host cell envelope, long flexible tail mechanism / viral tail assembly / virus tail / pore complex / monoatomic ion transport / cell outer membrane / host cell cytoplasm / entry receptor-mediated virion attachment to host cell ...maltodextrin transmembrane transporter activity / maltose transporting porin activity / symbiont genome ejection through host cell envelope, long flexible tail mechanism / viral tail assembly / virus tail / pore complex / monoatomic ion transport / cell outer membrane / host cell cytoplasm / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / virion attachment to host cell Similarity search - Function | |||||||||
| Biological species |  Shigella sonnei (bacteria) / Shigella sonnei (bacteria) /  Escherichia phage Lambda (virus) Escherichia phage Lambda (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.98 Å | |||||||||
 Authors Authors | Ge XF / Wang JW | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural mechanism of bacteriophage lambda tail's interaction with the bacterial receptor. Authors: Xiaofei Ge / Jiawei Wang /  Abstract: Bacteriophage infection, a pivotal process in microbiology, initiates with the phage's tail recognizing and binding to the bacterial cell surface, which then mediates the injection of viral DNA. ...Bacteriophage infection, a pivotal process in microbiology, initiates with the phage's tail recognizing and binding to the bacterial cell surface, which then mediates the injection of viral DNA. Although comprehensive studies on the interaction between bacteriophage lambda and its outer membrane receptor, LamB, have provided rich information about the system's biochemical properties, the precise molecular mechanism remains undetermined. This study revealed the high-resolution cryo-electron microscopy (cryo-EM) structures of the bacteriophage lambda tail complexed with its irreversible Shigella sonnei 3070 LamB receptor and the closed central tail fiber. These structures reveal the complex processes that trigger infection and demonstrate a substantial conformational change in the phage lambda tail tip upon LamB binding. Providing detailed structures of bacteriophage lambda infection initiation, this study contributes to the expanding knowledge of lambda-bacterial interaction, which holds significance in the fields of microbiology and therapeutic development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38245.map.gz emd_38245.map.gz | 168 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38245-v30.xml emd-38245-v30.xml emd-38245.xml emd-38245.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
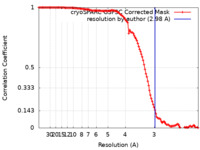
| FSC (resolution estimation) |  emd_38245_fsc.xml emd_38245_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_38245.png emd_38245.png | 116.7 KB | ||
| Filedesc metadata |  emd-38245.cif.gz emd-38245.cif.gz | 6.3 KB | ||
| Others |  emd_38245_half_map_1.map.gz emd_38245_half_map_1.map.gz emd_38245_half_map_2.map.gz emd_38245_half_map_2.map.gz | 165.3 MB 165.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38245 http://ftp.pdbj.org/pub/emdb/structures/EMD-38245 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38245 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38245 | HTTPS FTP |
-Related structure data
| Related structure data |  8xcjMC  8xcgC  8xciC  8xckC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38245.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38245.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0742 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_38245_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
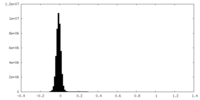
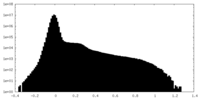
| Density Histograms |
-Half map: #2
| File | emd_38245_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
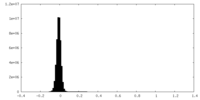
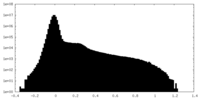
| Density Histograms |
- Sample components
Sample components
-Entire : Bacteriophage lambda tail with LamB
| Entire | Name: Bacteriophage lambda tail with LamB |
|---|---|
| Components |
|
-Supramolecule #1: Bacteriophage lambda tail with LamB
| Supramolecule | Name: Bacteriophage lambda tail with LamB / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: LamB
| Supramolecule | Name: LamB / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Shigella sonnei (bacteria) Shigella sonnei (bacteria) |
-Supramolecule #3: Tip attachment protein J
| Supramolecule | Name: Tip attachment protein J / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Escherichia phage Lambda (virus) Escherichia phage Lambda (virus) |
-Macromolecule #1: Maltoporin
| Macromolecule | Name: Maltoporin / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Shigella sonnei (bacteria) Shigella sonnei (bacteria) |
| Molecular weight | Theoretical: 47.568004 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AVDFHGYARS GIGWTGSGGE QQCFQTTGAQ SKYRLGNECE TYAELKLGQE VWKEGDKSFY FDTNVAYSVA QQNDWEATDP AFREANVQG KNLIEWLPGS TIWAGKRFYQ RHDVHMIDFY YWDISGPGAG LENIDVGFGK LSLAATRSSE AGGSSSFASN N IYDYTNET ...String: AVDFHGYARS GIGWTGSGGE QQCFQTTGAQ SKYRLGNECE TYAELKLGQE VWKEGDKSFY FDTNVAYSVA QQNDWEATDP AFREANVQG KNLIEWLPGS TIWAGKRFYQ RHDVHMIDFY YWDISGPGAG LENIDVGFGK LSLAATRSSE AGGSSSFASN N IYDYTNET ANDVFDVRLA QMEINPGGTL ELGVDYGRAN LRDNYRLVDG ASKDGWLFTA EHTQSVLKGF NKFVVQYATD SM TSQGKGL SQGSGVAFDN EKFAYNINNN GHMLRILDHG AISMGDNWDM MYVGMYQDIN WDNDNGTKWW TVGIRPMYKW TPI MSTVME IGYDNVESQR TGDKNNQYKI TLAQQWQAGD SIWSRPAIRV FATYAKWDEK WGYDYNGDSK VNPNYGKAVP ADFN GGSFG RGDSDEWTFG AQMEIWW UniProtKB: Maltoporin |
-Macromolecule #2: Tip attachment protein J
| Macromolecule | Name: Tip attachment protein J / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage Lambda (virus) Escherichia phage Lambda (virus) |
| Molecular weight | Theoretical: 46.045844 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: APAAPSRIEL TPGYFQITAT PHLAVYDPTV QFEFWFSEKQ IADIRQVETS TRYLGTALYW IAASINIKPG HDYYFYIRSV NTVGKSAFV EAVGRASDDA EGYLDFFKGK ITESHLGKEL LEKVELTEDN ASRLEEFSKE WKDASDKWNA MWAVKIEQTK D GKHYVAGI ...String: APAAPSRIEL TPGYFQITAT PHLAVYDPTV QFEFWFSEKQ IADIRQVETS TRYLGTALYW IAASINIKPG HDYYFYIRSV NTVGKSAFV EAVGRASDDA EGYLDFFKGK ITESHLGKEL LEKVELTEDN ASRLEEFSKE WKDASDKWNA MWAVKIEQTK D GKHYVAGI GLSMEDTEEG KLSQFLVAAN RIAFIDPANG NETPMFVAQG NQIFMNDVFL KRLTAPTITS GGNPPAFSLT PD GKLTAKN ADISGSVNAN SGTLSNVTIA ENCTINGTLR AEKIVGDIVK AASAAFPRQR ESSVDWPSGT RTVTVTDDHP FDR QIVVLP LTFRGSKRTV SGRTTYSMCY LKVLMNGAVI YDGAANEAVQ VFSRIVDMPA GRGNVILTFT LTSTRHSADI PPYT FASDV QVMVIKKQAL GISVV UniProtKB: Tip attachment protein J |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)