[English] 日本語
 Yorodumi
Yorodumi- EMDB-38087: Cryo-EM structure of Staphylococcus aureus sigA-dependent RNAP-pr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Staphylococcus aureus sigA-dependent RNAP-promoter open complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA polymerase / transcription / Staphylococcus aureus / sigA / TRANSCRIPTION-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsigma factor activity / DNA-directed RNA polymerase complex / DNA-templated transcription initiation / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / protein dimerization activity / DNA-templated transcription / magnesium ion binding / DNA binding ...sigma factor activity / DNA-directed RNA polymerase complex / DNA-templated transcription initiation / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / protein dimerization activity / DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Yuan L / Xu L / Liu Q / Feng Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis of promoter recognition by Staphylococcus aureus RNA polymerase. Authors: Linggang Yuan / Qingyang Liu / Liqiao Xu / Bing Wu / Yu Feng /  Abstract: Bacterial RNAP needs to form holoenzyme with σ factors to initiate transcription. While Staphylococcus aureus σ controls housekeeping functions, S. aureus σ regulates virulence, biofilm formation, ...Bacterial RNAP needs to form holoenzyme with σ factors to initiate transcription. While Staphylococcus aureus σ controls housekeeping functions, S. aureus σ regulates virulence, biofilm formation, persistence, cell internalization, membrane transport, and antimicrobial resistance. Besides the sequence difference, the spacers between the -35 element and -10 element of σ regulated promoters are shorter than those of σ regulated promoters. Therefore, how σ recognizes and initiates transcription from target promoters can not be inferred from that of the well studied σ. Here, we report the cryo-EM structures of S. aureus RNAP-promoter open complexes comprising σ and σ, respectively. Structural analyses, in combination with biochemical experiments, reveal the structural basis for the promoter specificity of S. aureus transcription. Although the -10 element of σ regulated promoters is recognized by domain σ as single-stranded DNA, the -10 element of σ regulated promoters is co-recognized by domains σ and σ as double-stranded DNA, accounting for the short spacers of σ regulated promoters. S. aureus RNAP is a validated target of antibiotics, and our structures pave the way for rational drug design targeting S. aureus RNAP. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38087.map.gz emd_38087.map.gz | 37.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38087-v30.xml emd-38087-v30.xml emd-38087.xml emd-38087.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_38087.png emd_38087.png | 155.5 KB | ||
| Filedesc metadata |  emd-38087.cif.gz emd-38087.cif.gz | 7.9 KB | ||
| Others |  emd_38087_half_map_1.map.gz emd_38087_half_map_1.map.gz emd_38087_half_map_2.map.gz emd_38087_half_map_2.map.gz | 31.3 MB 31.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38087 http://ftp.pdbj.org/pub/emdb/structures/EMD-38087 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38087 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38087 | HTTPS FTP |
-Related structure data
| Related structure data |  8x6fMC  8x6gC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38087.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38087.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.19 Å | ||||||||||||||||||||||||||||||||||||
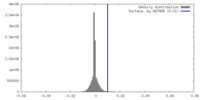
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_38087_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
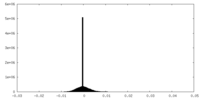
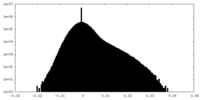
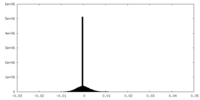
| Density Histograms |
-Half map: #1
| File | emd_38087_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : sigA-dependent RNAP-promoter open complex
| Entire | Name: sigA-dependent RNAP-promoter open complex |
|---|---|
| Components |
|
-Supramolecule #1: sigA-dependent RNAP-promoter open complex
| Supramolecule | Name: sigA-dependent RNAP-promoter open complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: DNA-directed RNA polymerase subunit alpha
| Macromolecule | Name: DNA-directed RNA polymerase subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 35.052656 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIEIEKPRIE TIEISEDAKF GKFVVEPLER GYGTTLGNSL RRILLSSLPG AAVKYIEIEG VLHEFSAVDN VVEDVSTIIM NIKQLALKI YSEEDKTLEI DVRDEGEVTA SDITHDSDVE ILNPELKIAT VSKGGHLKIR LVANKGRGYA LAEQNNTSDL P IGVIPVDS ...String: MIEIEKPRIE TIEISEDAKF GKFVVEPLER GYGTTLGNSL RRILLSSLPG AAVKYIEIEG VLHEFSAVDN VVEDVSTIIM NIKQLALKI YSEEDKTLEI DVRDEGEVTA SDITHDSDVE ILNPELKIAT VSKGGHLKIR LVANKGRGYA LAEQNNTSDL P IGVIPVDS LYSPVERVNY TVENTRVGQS SDFDKLTLDV WTNGSITPQE SVSLAAKIMT EHLNIFVGLT DEAQNAEIMI EK EEDQKEK VLEMSIEELD LSVRSYNCLK RAGINSVQEL ADKSEADMMK VRNLGRKSLE EVKYKLEDLG LGLRKED UniProtKB: DNA-directed RNA polymerase subunit alpha |
-Macromolecule #2: DNA-directed RNA polymerase subunit beta
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 133.381312 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGQVVQYGR HRKRRNYARI SEVLELPNLI EIQTKSYEWF LREGLIEMFR DISPIEDFTG NLSLEFVDYR LGEPKYDLEE SKNRDATYA APLRVKVRLI IKETGEVKEQ EVFMGDFPLM TDTGTFVING AERVIVSQLV RSPSVYFNEK IDKNGRENYD A TIIPNRGA ...String: MAGQVVQYGR HRKRRNYARI SEVLELPNLI EIQTKSYEWF LREGLIEMFR DISPIEDFTG NLSLEFVDYR LGEPKYDLEE SKNRDATYA APLRVKVRLI IKETGEVKEQ EVFMGDFPLM TDTGTFVING AERVIVSQLV RSPSVYFNEK IDKNGRENYD A TIIPNRGA WLEYETDAKD VVYVRIDRTR KLPLTVLLRA LGFSSDQEIV DLLGDNEYLR NTLEKDGTEN TEQALLEIYE RL RPGEPPT VENAKSLLYS RFFDPKRYDL ASVGRYKTNK KLHLKHRLFN QKLAEPIVNT ETGEIVVEEG TVLDRRKIDE IMD VLESNA NSEVFELHGS VIDEPVEIQS IKVYVPNDDE GRTTTVIGNA FPDSEVKCIT PADIIASMSY FFNLLSGIGY TDDI DHLGN RRLRSVGELL QNQFRIGLSR MERVVRERMS IQDTESITPQ QLINIRPVIA SIKEFFGSSQ LSQFMDQANP LAELT HKRR LSALGPGGLT RERAQMEVRD VHYSHYGRMC PIETPEGPNI GLINSLSSYA RVNEFGFIET PYRKVDLDTH AITDQI DYL TADEEDSYVV AQANSKLDEN GRFMDDEVVC RFRGNNTVMA KEKMDYMDVS PKQVVSAATA CIPFLENDDS NRALMGA NM QRQAVPLMNP EAPFVGTGME HVAARDSGAA ITAKHRGRVE HVESNEILVR RLVEENGVEH EGELDRYPLA KFKRSNSG T CYNQRPIVAV GDVVEYNEIL ADGPSMELGE MALGRNVVVG FMTWDGYNYE DAVIMSERLV KDDVYTSIHI EEYESEARD TKLGPEEITR DIPNVSESAL KNLDDRGIVY IGAEVKDGDI LVGKVTPKGV TELTAEERLL HAIFGEKARE VRDTSLRVPH GAGGIVLDV KVFNREEGDD TLSPGVNQLV RVYIVQKRKI HVGDKMCGRH GNKGVISKIV PEEDMPYLPD GRPIDIMLNP L GVPSRMNI GQVLELHLGM AAKNLGIHVA SPVFDGANDD DVWSTIEEAG MARDGKTVLY DGRTGEPFDN RISVGVMYML KL AHMVDDK LHARSTGPYS LVTQQPLGGK AQFGGQRFGE MEVWALEAYG AAYTLQEILT YKSDDTVGRV KTYEAIVKGE NIS RPSVPE SFRVLMKELQ SLGLDVKVMD EQDNEIEMTD VDDDDVVERK VDLQQNDAPE TQKEVTD UniProtKB: DNA-directed RNA polymerase subunit beta |
-Macromolecule #3: DNA-directed RNA polymerase subunit beta'
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta' / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 135.594266 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIDVNNFHYM KIGLASPEKI RSWSFGEVKK PETINYRTLK PEKDGLFCER IFGPTKDWEC SCGKYKRVRY KGMVCDRCGV EVTKSKVRR ERMGHIELAA PVSHIWYFKG IPSRMGLLLD MSPRALEEVI YFASYVVVDP GPTGLEKKTL LSEAEFRDYY D KYPGQFVA ...String: MIDVNNFHYM KIGLASPEKI RSWSFGEVKK PETINYRTLK PEKDGLFCER IFGPTKDWEC SCGKYKRVRY KGMVCDRCGV EVTKSKVRR ERMGHIELAA PVSHIWYFKG IPSRMGLLLD MSPRALEEVI YFASYVVVDP GPTGLEKKTL LSEAEFRDYY D KYPGQFVA KMGAEGIKDL LEEIDLDEEL KLLRDELESA TGQRLTRAIK RLEVVESFRN SGNKPSWMIL DVLPIIPPEI RP MVQLDGG RFATSDLNDL YRRVINRNNR LKRLLDLGAP GIIVQNEKRM LQEAVDALID NGRRGRPVTG PGNRPLKSLS HML KGKQGR FRQNLLGKRV DYSGRSVIAV GPSLKMYQCG LPKEMALELF KPFVMKELVQ REIATNIKNA KSKIERMDDE VWDV LEEVI REHPVLLNRA PTLHRLGIQA FEPTLVEGRA IRLHPLVTTA YNADFDGDQM AVHVPLSKEA QAEARMLMLA AQNIL NPKD GKPVVTPSQD MVLGNYYLTL ERKDAVNTGA IFNNTNEVLK AYANGFVHLH TRIGVHASSF NNPTFTEEQN KKILAT SVG KIIFNEIIPD SFAYINEPTQ ENLERKTPNR YFIDPTTLGE GGLKEYFENE ELIEPFNKKF LGNIIAEVFN RFSITDT SM MLDRMKDLGF KFSSKAGITV GVADIVVLPD KQQILDEHEK LVDRITKQFN RGLITEEERY NAVVEIWTDA KDQIQGEL M QSLDKTNPIF MMSDSGARGN ASNFTQLAGM RGLMAAPSGK IIELPITSSF REGLTVLEYF ISTHGARKGL ADTALKTAD SGYLTRRLVD VAQDVIVREE DCGTDRGLLV SDIKEGTEMI EPFIERIEGR YSKETIRHPE TDEIIIRPDE LITPEIAKKI TDAGIEQMY IRSAFTCNAR HGVCEKCYGK NLATGEKVEV GEAVGTIAAQ SIGEPGTQLT MRTFHTGGVA GSDITQGLPR I QEIFEARN PKGQAVITEI EGVVEDIKLA KDRQQEIVVK GANETRSYLA SGTSRIIVEI GQPVQRGEVL TEGSIEPKNY LS VAGLNAT ESYLLKEVQK VYRMQGVEID DKHVEVMVRQ MLRKVRIIEA GDTKLLPGSL VDIHNFTDAN REAFKHRKRP ATA KPVLLG ITKASLETES FLSAASFQET TRVLTDAAIK GKRDDLLGLK ENVIIGKLIP AGTGMRRYSD VKYEKTAKPV AEVE SQTEV TE UniProtKB: DNA-directed RNA polymerase subunit beta' |
-Macromolecule #4: DNA-directed RNA polymerase subunit omega
| Macromolecule | Name: DNA-directed RNA polymerase subunit omega / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.161236 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLNPPLNQLT SQIKSKYLIA TTAAKRAREI DEQPETELLS EYHSFKPVGR ALEEIADGKI RPVISSDYYG KE UniProtKB: DNA-directed RNA polymerase subunit omega |
-Macromolecule #5: DNA-directed RNA polymerase subunit epsilon
| Macromolecule | Name: DNA-directed RNA polymerase subunit epsilon / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.764739 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAVFKVFYQH NRDEVIVREN TQSLYVEAQT EEQVRRYLKD RNFNIEFITK LEGAHLDYEK ENSEHFNVEI AK UniProtKB: DNA-directed RNA polymerase subunit epsilon |
-Macromolecule #6: RNA polymerase sigma factor SigA
| Macromolecule | Name: RNA polymerase sigma factor SigA / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.218754 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSDNTVKIKK QTIDPTLTLE DVKKQLIEKG KKEGHLSHEE IAEKLQNFDI DSDQMDDFFD QLNDNDISLV NEKDSSDTDE KLNPSDLSA PPGVKINDPV RMYLKEIGRV NLLSAQEEIE LAKRIEQGDE VAKSRLAEAN LRLVVSIAKR YVGRGMLFLD L IQEGNMGL ...String: MSDNTVKIKK QTIDPTLTLE DVKKQLIEKG KKEGHLSHEE IAEKLQNFDI DSDQMDDFFD QLNDNDISLV NEKDSSDTDE KLNPSDLSA PPGVKINDPV RMYLKEIGRV NLLSAQEEIE LAKRIEQGDE VAKSRLAEAN LRLVVSIAKR YVGRGMLFLD L IQEGNMGL IKAVEKFDFN KGFKFSTYAT WWIRQAITRA IADQARTIRI PVHMVETINK LIRVQRQLLQ DLGRDPAPEE IG EEMDLPA EKVREVLKIA QEPVSLETPI GEEDDSHLGD FIEDQEAQSP SDHAAYELLK EQLEDVLDTL TDREENVLRL RFG LDDGRT RTLEEVGKVF GVTRERIRQI EAKALRKLRH PSRSKRLKDF MD UniProtKB: RNA polymerase sigma factor SigA |
-Macromolecule #7: DNA (71-mer)
| Macromolecule | Name: DNA (71-mer) / type: dna / ID: 7 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.923221 KDa |
| Sequence | String: (DA)(DA)(DA)(DA)(DT)(DA)(DA)(DT)(DT)(DA) (DA)(DA)(DA)(DA)(DT)(DA)(DA)(DT)(DT)(DC) (DT)(DT)(DG)(DA)(DC)(DA)(DT)(DA)(DC) (DA)(DA)(DA)(DA)(DA)(DC)(DT)(DT)(DA)(DC) (DG) (DA)(DG)(DT)(DT)(DA)(DT) ...String: (DA)(DA)(DA)(DA)(DT)(DA)(DA)(DT)(DT)(DA) (DA)(DA)(DA)(DA)(DT)(DA)(DA)(DT)(DT)(DC) (DT)(DT)(DG)(DA)(DC)(DA)(DT)(DA)(DC) (DA)(DA)(DA)(DA)(DA)(DC)(DT)(DT)(DA)(DC) (DG) (DA)(DG)(DT)(DT)(DA)(DT)(DA)(DA) (DT)(DT)(DA)(DA)(DA)(DT)(DC)(DT)(DT)(DG) (DT)(DA) (DA)(DG)(DT)(DG)(DA)(DC)(DA) (DA)(DA)(DC)(DG) |
-Macromolecule #8: DNA (71-mer)
| Macromolecule | Name: DNA (71-mer) / type: dna / ID: 8 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.797018 KDa |
| Sequence | String: (DC)(DG)(DT)(DT)(DT)(DG)(DT)(DC)(DA)(DC) (DT)(DT)(DA)(DC)(DT)(DT)(DC)(DT)(DA)(DA) (DA)(DT)(DT)(DA)(DA)(DT)(DA)(DA)(DA) (DC)(DT)(DC)(DG)(DT)(DA)(DA)(DG)(DT)(DT) (DT) (DT)(DT)(DG)(DT)(DA)(DT) ...String: (DC)(DG)(DT)(DT)(DT)(DG)(DT)(DC)(DA)(DC) (DT)(DT)(DA)(DC)(DT)(DT)(DC)(DT)(DA)(DA) (DA)(DT)(DT)(DA)(DA)(DT)(DA)(DA)(DA) (DC)(DT)(DC)(DG)(DT)(DA)(DA)(DG)(DT)(DT) (DT) (DT)(DT)(DG)(DT)(DA)(DT)(DG)(DT) (DC)(DA)(DA)(DG)(DA)(DA)(DT)(DT)(DA)(DT) (DT)(DT) (DT)(DT)(DA)(DA)(DT)(DT)(DA) (DT)(DT)(DT)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 51.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.7 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 56457 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)