[English] 日本語
 Yorodumi
Yorodumi- EMDB-36284: Cryo-EM structure of human S1P transporter SPNS2 bound with S1P -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human S1P transporter SPNS2 bound with S1P | |||||||||
 Map data Map data | SPNS2 fusion full map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Transpoter / LIPID TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of eye pigmentation / regulation of humoral immune response / regulation of T cell migration / alpha-1,4-glucan glucosyltransferase (UDP-glucose donor) activity / sphingolipid intramembrane carrier activity / lymphocyte migration / sphingolipid biosynthetic process / Sphingolipid de novo biosynthesis / sphingosine-1-phosphate receptor signaling pathway / lipid transport ...regulation of eye pigmentation / regulation of humoral immune response / regulation of T cell migration / alpha-1,4-glucan glucosyltransferase (UDP-glucose donor) activity / sphingolipid intramembrane carrier activity / lymphocyte migration / sphingolipid biosynthetic process / Sphingolipid de novo biosynthesis / sphingosine-1-phosphate receptor signaling pathway / lipid transport / B cell homeostasis / T cell homeostasis / lymph node development / transmembrane transporter activity / sensory perception of sound / bone development / endosome membrane / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Pang B / Yu LY / Ren RB | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2024 Journal: Cell Res / Year: 2024Title: Molecular basis of Spns2-facilitated sphingosine-1-phosphate transport. Authors: Bin Pang / Leiye Yu / Tong Li / Haizhan Jiao / Xiaomei Wu / Jinxin Wang / Ruiping He / Yurou Zhang / Juan Wang / Hongli Hu / Wei Dai / Li Chen / Ruobing Ren /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36284.map.gz emd_36284.map.gz | 85.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36284-v30.xml emd-36284-v30.xml emd-36284.xml emd-36284.xml | 14.5 KB 14.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36284.png emd_36284.png | 41.3 KB | ||
| Filedesc metadata |  emd-36284.cif.gz emd-36284.cif.gz | 5.8 KB | ||
| Others |  emd_36284_half_map_1.map.gz emd_36284_half_map_1.map.gz emd_36284_half_map_2.map.gz emd_36284_half_map_2.map.gz | 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36284 http://ftp.pdbj.org/pub/emdb/structures/EMD-36284 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36284 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36284 | HTTPS FTP |
-Related structure data
| Related structure data |  8jhqMC  8jhrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36284.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36284.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SPNS2 fusion full map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
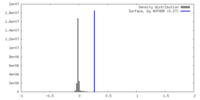
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: SPNS2 fusion half B map
| File | emd_36284_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SPNS2 fusion half B map | ||||||||||||
| Projections & Slices |
| ||||||||||||
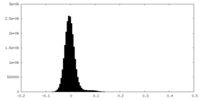
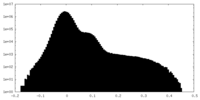
| Density Histograms |
-Half map: SPNS2 fusion half A map
| File | emd_36284_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SPNS2 fusion half A map | ||||||||||||
| Projections & Slices |
| ||||||||||||
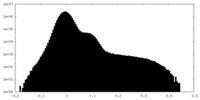
| Density Histograms |
- Sample components
Sample components
-Entire : SPNS2 fused with PGS
| Entire | Name: SPNS2 fused with PGS |
|---|---|
| Components |
|
-Supramolecule #1: SPNS2 fused with PGS
| Supramolecule | Name: SPNS2 fused with PGS / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Sphingosine-1-phosphate transporter SPNS2,GlgA glycogen synthase
| Macromolecule | Name: Sphingosine-1-phosphate transporter SPNS2,GlgA glycogen synthase type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 82.08993 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDDDDKL EVLFQGPGSM MCLECASAAA GGAEEEEADA ERRRRRRGAQ RGAGGSGCCG ARGAGGAGVS AAGDEVQTLS GSVRRAPTG PPGTPGTPGC AATAKGPGAQ QPKPASLGRG RGAAAAILSL GNVLNYLDRY TVAGVLLDIQ QHFGVKDRGA G LLQSVFIC ...String: MDYKDDDDKL EVLFQGPGSM MCLECASAAA GGAEEEEADA ERRRRRRGAQ RGAGGSGCCG ARGAGGAGVS AAGDEVQTLS GSVRRAPTG PPGTPGTPGC AATAKGPGAQ QPKPASLGRG RGAAAAILSL GNVLNYLDRY TVAGVLLDIQ QHFGVKDRGA G LLQSVFIC SFMVAAPIFG YLGDRFNRKV ILSCGIFFWS AVTFSSSFIP QQYFWLLVLS RGLVGIGEAS YSTIAPTIIG DL FGIDCSF WNESYLTGSR DERKKSLLSK FGMDEGVTFM FIGRFDRGQK GVDVLLKAIE ILSSKKEFQE MRFIIIGKGD PEL EGWARS LEEKHGNVKV ITEMLSREFV RELYGSVDFV IIPSYFEPFG LVALEAMCLG AIPIASAVGG LRDIITNETG ILVK AGDPG ELANAILKAL ELSRSDLSKF RENCKKRAMS FSTKNTRTLM LSVFYFAIPL GSGLGYITGS SVKQAAGDWH WALRV SPVL GMITGTLILI LVPATKRGHA DQLGDQLKAR TSWLRDMKAL IRNRSYVFSS LATSAVSFAT GALGMWIPLY LHRAQV VQK TAETCNSPPC GAKDSLIFGA ITCFTGFLGV VTGAGATRWC RLKTQRADPL VCAVGMLGSA IFICLIFVAA KSSIVGA YI CIFVGETLLF SNWAITADIL MYVVIPTRRA TAVALQSFTS HLLGDAGSPY LIGFISDLIR QSTKDSPLWE FLSLGYAL M LCPFVVVLGG MFFLATALFF VSDRARAEQQ VNQLAMPPAS VKV UniProtKB: Sphingosine-1-phosphate transporter SPNS2, Glycogen synthase, Sphingosine-1-phosphate transporter SPNS2 |
-Macromolecule #2: (2S,3R,4E)-2-amino-3-hydroxyoctadec-4-en-1-yl dihydrogen phosphate
| Macromolecule | Name: (2S,3R,4E)-2-amino-3-hydroxyoctadec-4-en-1-yl dihydrogen phosphate type: ligand / ID: 2 / Number of copies: 1 / Formula: S1P |
|---|---|
| Molecular weight | Theoretical: 379.472 Da |
| Chemical component information | 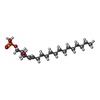 ChemComp-S1P: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 54.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 390253 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)