+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Na+,K+-ATPase in the E1.Mg2+ state. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  ion pump / ion pump /  P-type ATPase / P-type ATPase /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of monoatomic ion transport / P-type potassium transmembrane transporter activity / sodium:potassium-exchanging ATPase complex / ion channel regulator activity / sodium ion transport / monoatomic ion transport / potassium ion transport /  ATP hydrolysis activity / ATP hydrolysis activity /  ATP binding / ATP binding /  membrane ...regulation of monoatomic ion transport / P-type potassium transmembrane transporter activity / sodium:potassium-exchanging ATPase complex / ion channel regulator activity / sodium ion transport / monoatomic ion transport / potassium ion transport / membrane ...regulation of monoatomic ion transport / P-type potassium transmembrane transporter activity / sodium:potassium-exchanging ATPase complex / ion channel regulator activity / sodium ion transport / monoatomic ion transport / potassium ion transport /  ATP hydrolysis activity / ATP hydrolysis activity /  ATP binding / ATP binding /  membrane / membrane /  metal ion binding / metal ion binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Squalus acanthias (spiny dogfish) Squalus acanthias (spiny dogfish) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.5 Å cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Kanai R / Vilsen B / Cornelius F / Toyoshima C | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: FEBS Lett / Year: 2023 Journal: FEBS Lett / Year: 2023Title: Crystal structures of Na ,K -ATPase reveal the mechanism that converts the K -bound form to Na -bound form and opens and closes the cytoplasmic gate. Authors: Ryuta Kanai / Bente Vilsen / Flemming Cornelius / Chikashi Toyoshima /   Abstract: Na ,K -ATPase (NKA) plays a pivotal role in establishing electrochemical gradients for Na and K across the cell membrane by alternating between the E1 (showing high affinity for Na and low affinity ...Na ,K -ATPase (NKA) plays a pivotal role in establishing electrochemical gradients for Na and K across the cell membrane by alternating between the E1 (showing high affinity for Na and low affinity for K ) and E2 (low affinity to Na and high affinity to K ) forms. Presented here are two crystal structures of NKA in E1·Mg and E1·3Na states at 2.9 and 2.8 Å resolution, respectively. These two E1 structures fill a gap in our description of the NKA reaction cycle based on the atomic structures. We describe how NKA converts the K -bound E2·2K form to an E1 (E1·Mg ) form, which allows high-affinity Na binding, eventually closing the cytoplasmic gate (in E1 ~ P·ADP·3Na ) after binding three Na , while keeping the extracellular ion pathway sealed. We now understand previously unknown functional roles for several parts of NKA and that NKA uses even the lipid bilayer for gating the ion pathway. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36220.map.gz emd_36220.map.gz | 48.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36220-v30.xml emd-36220-v30.xml emd-36220.xml emd-36220.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
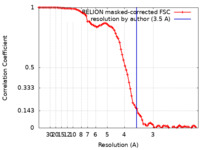
| FSC (resolution estimation) |  emd_36220_fsc.xml emd_36220_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_36220.png emd_36220.png | 88.9 KB | ||
| Others |  emd_36220_half_map_1.map.gz emd_36220_half_map_1.map.gz emd_36220_half_map_2.map.gz emd_36220_half_map_2.map.gz | 40.5 MB 40.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36220 http://ftp.pdbj.org/pub/emdb/structures/EMD-36220 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36220 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36220 | HTTPS FTP |
-Related structure data
| Related structure data |  8jfzMC  8jbkC  8jblC  8jbmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36220.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36220.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.076 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36220_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36220_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Na+,K+-ATPase
| Entire | Name: Na+,K+-ATPase |
|---|---|
| Components |
|
-Supramolecule #1: Na+,K+-ATPase
| Supramolecule | Name: Na+,K+-ATPase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:   Squalus acanthias (spiny dogfish) Squalus acanthias (spiny dogfish) |
-Macromolecule #1: Na+,K+-ATPase beta subunit
| Macromolecule | Name: Na+,K+-ATPase beta subunit / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Squalus acanthias (spiny dogfish) Squalus acanthias (spiny dogfish) |
| Molecular weight | Theoretical: 35.176125 KDa |
| Sequence | String: MARGKSKETD GGWKKFLWDS EKKEFLGRTG SSWFKIFLFY LIFYGCLAGI FIGTIQVLLL TLSDFEPKYQ DRVAPPGLSH APYAIKTEI SFSISNPKSY ESFVKSMHKL MDLYNESSQA GNSPFEDCSD TPADYIKRGD LDDSQGQKKA CRFSRMWLKN C SGLDDTTY ...String: MARGKSKETD GGWKKFLWDS EKKEFLGRTG SSWFKIFLFY LIFYGCLAGI FIGTIQVLLL TLSDFEPKYQ DRVAPPGLSH APYAIKTEI SFSISNPKSY ESFVKSMHKL MDLYNESSQA GNSPFEDCSD TPADYIKRGD LDDSQGQKKA CRFSRMWLKN C SGLDDTTY GYAEGKPCVV AKLNRIIGFY PKPLKNTTDL PEELQANYNQ YVLPLRCAAK REEDREKIGS IEYFGLGGYA GF PLQYYPY YGKRLQKKYL QPLLAIQFTN LTQNMELRIE CKVYGENIDY SEKDRFRGRF EVKIEVKS UniProtKB: Sodium/potassium-transporting ATPase subunit beta |
-Macromolecule #2: Na, K-ATPase alpha subunit
| Macromolecule | Name: Na, K-ATPase alpha subunit / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Squalus acanthias (spiny dogfish) Squalus acanthias (spiny dogfish) |
| Molecular weight | Theoretical: 113.309891 KDa |
| Sequence | String: MGKGTASDKY EPAATSENAT KSKKKGKKDK IDKKRDLDEL KKEVSMDDHK LSLDELHNKY GTDLTRGLTN ARAKEILARD GPNSLTPPP TTPEWIKFCR QLFGGFSILL WIGAILCFLA YGIQAATEDE PANDNLYLGV VLSTVVIVTG CFSYYQEAKS S RIMDSFKN ...String: MGKGTASDKY EPAATSENAT KSKKKGKKDK IDKKRDLDEL KKEVSMDDHK LSLDELHNKY GTDLTRGLTN ARAKEILARD GPNSLTPPP TTPEWIKFCR QLFGGFSILL WIGAILCFLA YGIQAATEDE PANDNLYLGV VLSTVVIVTG CFSYYQEAKS S RIMDSFKN MVPQQALVIR DGEKSTINAE FVVAGDLVEV KGGDRIPADL RIISAHGCKV DNSSLTGESE PQTRSPEFSS EN PLETRNI AFFSTNCVEG TARGVVVYTG DRTVMGRIAT LASGLEVGRT PIAIEIEHFI HIITGVAVFL GVSFFILSLI LGY SWLEAV IFLIGIIVAN VPEGLLATVT VCLTLTAKRM ARKNCLVKNL EAVETLGSTS TICSDKTGTL TQNRMTVAHM WFDN QIHEA DTTENQSGAA FDKTSATWSA LSRIAALCNR AVFQAGQDNV PILKRSVAGD ASESALLKCI ELCCGSVQGM RDRNP KIVE IPFNSTNKYQ LSIHENEKSS ESRYLLVMKG APERILDRCS TILLNGAEEP LKEDMKEAFQ NAYLELGGLG ERVLGF CHF ALPEDKYNEG YPFDADEPNF PTTDLCFVGL MAMIDPPRAA VPDAVGKCRS AGIKVIMVTG DHPITAKAIA KGVGIIS EG NETIEDIAAR LNIPIGQVNP RDAKACVVHG SDLKDLSTEV LDDILHYHTE IVFARTSPQQ KLIIVEGCQR QGAIVAVT G DGVNDSPALK KADIGVAMGI SGSDVSKQAA DMILLDDNFA SIVTGVEEGR LIFDNLKKSI AYTLTSNIPE ITPFLVFII GNVPLPLGTV TILCIDLGTD MVPAISLAYE QAESDIMKRQ PRNPKTDKLV NERLISMAYG QIGMIQALGG FFSYFVILAE NGFLPMDLI GKRVRWDDRW ISDVEDSFGQ QWTYEQRKIV EFTCHTSFFI SIVVVQWADL IICKTRRNSI FQQGMKNKIL I FGLFEETA LAAFLSYCPG TDVALRMYPL KPSWWFCAFP YSLIIFLYDE MRRFIIRRSP GGWVEQETYY UniProtKB: Sodium/potassium-transporting ATPase subunit alpha |
-Macromolecule #3: FXYD domain-containing ion transport regulator
| Macromolecule | Name: FXYD domain-containing ion transport regulator / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Squalus acanthias (spiny dogfish) Squalus acanthias (spiny dogfish) |
| Molecular weight | Theoretical: 10.195847 KDa |
| Sequence | String: MLGAATGLMV LVAVTQGVWA MDPEGPDNDE RFTYDYYRLR VVGLIVAAVL CVIGIIILLA GKCRCKFNQN KRTRSNSGTA TAQHLLQPG EATEC UniProtKB: FXYD domain-containing ion transport regulator |
-Macromolecule #8: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 8 / Number of copies: 8 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #9: 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 9 / Number of copies: 20 / Formula: PCW |
|---|---|
| Molecular weight | Theoretical: 787.121 Da |
| Chemical component information |  ChemComp-PCW: |
-Macromolecule #10: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 10 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #11: water
| Macromolecule | Name: water / type: ligand / ID: 11 / Number of copies: 6 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 99.9 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X