+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of human HCN3 channel with cilobradine | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | human HCN3 channel / MEMBRANE PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報HCN channels / HCN channel complex / regulation of SA node cell action potential / sodium ion import across plasma membrane / voltage-gated sodium channel activity / regulation of membrane depolarization / potassium ion import across plasma membrane / regulation of heart rate by cardiac conduction / voltage-gated potassium channel activity / cAMP binding ...HCN channels / HCN channel complex / regulation of SA node cell action potential / sodium ion import across plasma membrane / voltage-gated sodium channel activity / regulation of membrane depolarization / potassium ion import across plasma membrane / regulation of heart rate by cardiac conduction / voltage-gated potassium channel activity / cAMP binding / potassium ion transmembrane transport / cellular response to cAMP / sodium ion transmembrane transport / regulation of membrane potential / axon / dendrite / plasma membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.02 Å | |||||||||
 データ登録者 データ登録者 | Yu B / Lu QY / Li J / Zhang J | |||||||||
| 資金援助 | 1件
| |||||||||
 引用 引用 |  ジャーナル: To Be Published ジャーナル: To Be Publishedタイトル: Cryo-EM structure of human HCN3 channel with cilobradine 著者: Yu B / Lu QY / Li J / Zhang J | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_35606.map.gz emd_35606.map.gz | 112.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-35606-v30.xml emd-35606-v30.xml emd-35606.xml emd-35606.xml | 16.1 KB 16.1 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_35606.png emd_35606.png | 32.3 KB | ||
| Filedesc metadata |  emd-35606.cif.gz emd-35606.cif.gz | 6 KB | ||
| その他 |  emd_35606_half_map_1.map.gz emd_35606_half_map_1.map.gz emd_35606_half_map_2.map.gz emd_35606_half_map_2.map.gz | 224.2 MB 224.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35606 http://ftp.pdbj.org/pub/emdb/structures/EMD-35606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35606 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8io3MC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_35606.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_35606.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.526 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #2
| ファイル | emd_35606_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
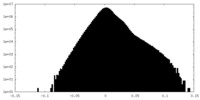
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_35606_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
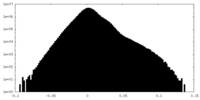
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : otassium/sodium hyperpolarization-activated cyclic nucleotide-gat...
| 全体 | 名称: otassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 |
|---|---|
| 要素 |
|
-超分子 #1: otassium/sodium hyperpolarization-activated cyclic nucleotide-gat...
| 超分子 | 名称: otassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: Potassium/sodium hyperpolarization-activated cyclic nucleotide-ga...
| 分子 | 名称: Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 タイプ: protein_or_peptide / ID: 1 / コピー数: 4 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 86.142922 KDa |
| 組換発現 | 生物種: Mammalian expression vector EGFP-MCS-pcDNA3.1 (その他) |
| 配列 | 文字列: MEAEQRPAAG ASEGATPGLE AVPPVAPPPA TAASGPIPKS GPEPKRRHLG TLLQPTVNKF SLRVFGSHKA VEIEQERVKS AGAWIIHPY SDFRFYWDLI MLLLMVGNLI VLPVGITFFK EENSPPWIVF NVLSDTFFLL DLVLNFRTGI VVEEGAEILL A PRAIRTRY ...文字列: MEAEQRPAAG ASEGATPGLE AVPPVAPPPA TAASGPIPKS GPEPKRRHLG TLLQPTVNKF SLRVFGSHKA VEIEQERVKS AGAWIIHPY SDFRFYWDLI MLLLMVGNLI VLPVGITFFK EENSPPWIVF NVLSDTFFLL DLVLNFRTGI VVEEGAEILL A PRAIRTRY LRTWFLVDLI SSIPVDYIFL VVELEPRLDA EVYKTARALR IVRFTKILSL LRLLRLSRLI RYIHQWEEIF HM TYDLASA VVRIFNLIGM MLLLCHWDGC LQFLVPMLQD FPPDCWVSIN HMVNHSWGRQ YSHALFKAMS HMLCIGYGQQ APV GMPDVW LTMLSMIVGA TCYAMFIGHA TALIQSLDSS RRQYQEKYKQ VEQYMSFHKL PADTRQRIHE YYEHRYQGKM FDEE SILGE LSEPLREEII NFTCRGLVAH MPLFAHADPS FVTAVLTKLR FEVFQPGDLV VREGSVGRKM YFIQHGLLSV LARGA RDTR LTDGSYFGEI CLLTRGRRTA SVRADTYCRL YSLSVDHFNA VLEEFPMMRR AFETVAMDRL LRIGKKNSIL QRKRSE PSP GSSGGIMEQH LVQHDRDMAR GVRGRAPSTG AQLSGKPVLW EPLVHAPLQA AAVTSNVAIA LTHQRGPLPL SPDSPAT LL ARSAWRSAGS PASPLVPVRA GPWASTSRLP APPARTLHAS LSRAGRSQVS LLGPPPGGGG RRLGPRGRPL SASQPSLP Q RATGDGSPGR KGSGSERLPP SGLLAKPPRT AQPPRPPVPE PATPRGLQLS ANM UniProtKB: Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 |
-分子 #2: 3-[[(3~{S})-1-[2-(3,4-dimethoxyphenyl)ethyl]piperidin-3-yl]methyl...
| 分子 | 名称: 3-[[(3~{S})-1-[2-(3,4-dimethoxyphenyl)ethyl]piperidin-3-yl]methyl]-7,8-dimethoxy-2,5-dihydro-1~{H}-3-benzazepin-4-one タイプ: ligand / ID: 2 / コピー数: 4 / 式: VXI |
|---|---|
| 分子量 | 理論値: 482.612 Da |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 40.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.2 µm / 最小 デフォーカス(公称値): 1.8 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)