[English] 日本語
 Yorodumi
Yorodumi- EMDB-34921: The cryo-EM structure of cellobiose phosphorylase from Clostridiu... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The cryo-EM structure of cellobiose phosphorylase from Clostridium thermocellum | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cellobiose phosphorylase / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationcellobiose phosphorylase / cellobiose phosphorylase activity / carbohydrate binding / carbohydrate metabolic process Similarity search - Function | |||||||||
| Biological species |  Acetivibrio thermocellus (bacteria) Acetivibrio thermocellus (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.59 Å | |||||||||
 Authors Authors | Iriya S / Kuga T / Sunagawa N / Igarashi K | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The cryo-EM structure of cellobiose phosphorylase from Clostridium thermocellum Authors: Iriya S / Kuga T / Sunagawa N / Igarashi K | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34921.map.gz emd_34921.map.gz | 225.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34921-v30.xml emd-34921-v30.xml emd-34921.xml emd-34921.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
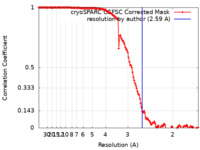
| FSC (resolution estimation) |  emd_34921_fsc.xml emd_34921_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_34921.png emd_34921.png | 89.2 KB | ||
| Masks |  emd_34921_msk_1.map emd_34921_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-34921.cif.gz emd-34921.cif.gz | 6 KB | ||
| Others |  emd_34921_half_map_1.map.gz emd_34921_half_map_1.map.gz emd_34921_half_map_2.map.gz emd_34921_half_map_2.map.gz | 226.2 MB 226.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34921 http://ftp.pdbj.org/pub/emdb/structures/EMD-34921 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34921 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34921 | HTTPS FTP |
-Related structure data
| Related structure data |  8ho7MC  8ho9C  8hobC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34921.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34921.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_34921_msk_1.map emd_34921_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34921_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_34921_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cellobiose phosphorylase
| Entire | Name: Cellobiose phosphorylase |
|---|---|
| Components |
|
-Supramolecule #1: Cellobiose phosphorylase
| Supramolecule | Name: Cellobiose phosphorylase / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Acetivibrio thermocellus (bacteria) / Strain: YM4 Acetivibrio thermocellus (bacteria) / Strain: YM4 |
| Molecular weight | Theoretical: 180 KDa |
-Macromolecule #1: Cellobiose phosphorylase
| Macromolecule | Name: Cellobiose phosphorylase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: cellobiose phosphorylase |
|---|---|
| Source (natural) | Organism:  Acetivibrio thermocellus (bacteria) Acetivibrio thermocellus (bacteria) |
| Molecular weight | Theoretical: 93.872453 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEFGFFDDAN KEYVITVPRT PYPWINYLGT ENFFSLISNT AGGYCFYRDA RLRRITRYRY NNVPIDMGGR YFYIYDNGDF WSPGWSPVK RELESYECRH GLGYTKIAGK RNGIKAEVTF FVPLNYNGEV QKLILKNEGQ DKKKITLFSF IEFCLWNAYD D MTNFQRNF ...String: MEFGFFDDAN KEYVITVPRT PYPWINYLGT ENFFSLISNT AGGYCFYRDA RLRRITRYRY NNVPIDMGGR YFYIYDNGDF WSPGWSPVK RELESYECRH GLGYTKIAGK RNGIKAEVTF FVPLNYNGEV QKLILKNEGQ DKKKITLFSF IEFCLWNAYD D MTNFQRNF STGEVEIEGS VIYHKTEYRE RRNHYAFYSV NAKISGFDSD RDSFIGLYNG FDAPQAVVNG KSNNSVADGW AP IASHSIE IELNPGEQKE YVFIIGYVEN KDEEKWESKG VINKKKAYEM IEQFNTVEKV DKAFEELKSY WNALLSKYFL ESH DEKLNR MVNIWNQYQC MVTFNMSRSA SYFESGIGRG MGFRDSNQDL LGFVHQIPER ARERLLDLAA TQLEDGSAYH QYQP LTKKG NNEIGSNFND DPLWLILATA AYIKETGDYS ILKEQVPFNN DPSKADTMFE HLTRSFYHVV NNLGPHGLPL IGRAD WNDC LNLNCFSTVP DESFQTTTSK DGKVAESVMI AGMFVFIGKD YVKLCEYMGL EEEARKAQQH IDAMKEAILK YGYDGE WFL RAYDDFGRKV GSKENEEGKI FIESQGFCVM AEIGLEDGKA LKALDSVKKY LDTPYGLVLQ NPAFTRYYIE YGEISTY PP GYKENAGIFC HNNAWIICAE TVVGRGDMAF DYYRKIAPAY IEDVSDIHKL EPYVYAQMVA GKDAKRHGEA KNSWLTGT A AWNFVAISQW ILGVKPDYDG LKIDPCIPKA WDGYKVTRYF RGSTYEITVK NPNHVSKGVA KITVDGNEIS GNILPVFND GKTHKVEVIL EHHHHHH UniProtKB: Cellobiose phosphorylase |
-Macromolecule #2: water
| Macromolecule | Name: water / type: ligand / ID: 2 / Number of copies: 112 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6.5 Component:
Details: 20 mM MES, 70 mM NaCl | |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV Details: Vitrification carried out in nitrogen atmosphere.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 2907 / Average exposure time: 5.121 sec. / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)