[English] 日本語
 Yorodumi
Yorodumi- EMDB-34073: Cryo-EM structure of the Serotonin 6 (5-HT6) receptor-DNGs-scFv16... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Serotonin 6 (5-HT6) receptor-DNGs-scFv16 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / serotonin receptor / 5-HT6R / cryo-EM / constitutive activity / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationhistamine receptor activity / Serotonin receptors / G protein-coupled serotonin receptor activity / neurotransmitter receptor activity / cerebral cortex cell migration / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / positive regulation of TOR signaling / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta ...histamine receptor activity / Serotonin receptors / G protein-coupled serotonin receptor activity / neurotransmitter receptor activity / cerebral cortex cell migration / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / positive regulation of TOR signaling / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Sensory perception of sweet, bitter, and umami (glutamate) taste / cilium / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade / heterotrimeric G-protein complex / G alpha (12/13) signalling events / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / GTPase binding / retina development in camera-type eye / phospholipase C-activating G protein-coupled receptor signaling pathway / Ca2+ pathway / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / G alpha (q) signalling events / chemical synaptic transmission / cell population proliferation / Ras protein signal transduction / Extra-nuclear estrogen signaling / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / synapse / dendrite / protein-containing complex binding / signal transduction / extracellular exosome / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Zhao QY / Wang YF / He L / Wang S / Cong Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structural insights into constitutive activity of 5-HT receptor. Authors: Licong He / Qiaoyu Zhao / Jianzhong Qi / Yifan Wang / Wenyu Han / Zhangcheng Chen / Yao Cong / Sheng Wang /  Abstract: While most therapeutic research on G-protein-coupled receptors (GPCRs) focuses on receptor activation by (endogenous) agonists, significant therapeutic potential exists through agonist-independent ...While most therapeutic research on G-protein-coupled receptors (GPCRs) focuses on receptor activation by (endogenous) agonists, significant therapeutic potential exists through agonist-independent intrinsic constitutive activity that can occur in various physiological and pathophysiological settings. For example, inhibiting the constitutive activity of 5-HTR-a receptor that is found almost exclusively in the brain and mediates excitatory neurotransmission-has demonstrated a therapeutic effect on cognitive/memory impairment associated with several neuropsychiatric disorders. However, the structural basis of such constitutive activity remains unclear. Here, we present a cryo-EM structure of serotonin-bound human 5-HTR-Gs heterotrimer at 3.0-Å resolution. Detailed analyses of the structure complemented by comprehensive interrogation of signaling illuminate key structural determinants essential for constitutive 5-HTR activity. Additional structure-guided mutagenesis leads to a nanobody mimic Gαs for 5-HTR that can reduce its constitutive activity. Given the importance of 5-HTR for a large number of neuropsychiatric disorders, insights derived from these studies will accelerate the design of more effective medications, and shed light on the molecular basis of constitutive activity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34073.map.gz emd_34073.map.gz | 56.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34073-v30.xml emd-34073-v30.xml emd-34073.xml emd-34073.xml | 19 KB 19 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34073.png emd_34073.png | 100 KB | ||
| Filedesc metadata |  emd-34073.cif.gz emd-34073.cif.gz | 6.3 KB | ||
| Others |  emd_34073_half_map_1.map.gz emd_34073_half_map_1.map.gz emd_34073_half_map_2.map.gz emd_34073_half_map_2.map.gz | 49.6 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34073 http://ftp.pdbj.org/pub/emdb/structures/EMD-34073 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34073 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34073 | HTTPS FTP |
-Validation report
| Summary document |  emd_34073_validation.pdf.gz emd_34073_validation.pdf.gz | 649 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34073_full_validation.pdf.gz emd_34073_full_validation.pdf.gz | 648.5 KB | Display | |
| Data in XML |  emd_34073_validation.xml.gz emd_34073_validation.xml.gz | 11.9 KB | Display | |
| Data in CIF |  emd_34073_validation.cif.gz emd_34073_validation.cif.gz | 14 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34073 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34073 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34073 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34073 | HTTPS FTP |
-Related structure data
| Related structure data |  7ys6MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34073.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34073.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.093 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_34073_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_34073_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Serotonin 6 (5-HT6) receptor-DNGs-scFv16 complex
| Entire | Name: Serotonin 6 (5-HT6) receptor-DNGs-scFv16 complex |
|---|---|
| Components |
|
-Supramolecule #1: Serotonin 6 (5-HT6) receptor-DNGs-scFv16 complex
| Supramolecule | Name: Serotonin 6 (5-HT6) receptor-DNGs-scFv16 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: 5-hydroxytryptamine receptor 6
| Macromolecule | Name: 5-hydroxytryptamine receptor 6 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 33.201918 KDa |
| Recombinant expression | Organism:  Insecta environmental sample (insect) Insecta environmental sample (insect) |
| Sequence | String: GGSGWVAAAL CVVIALTAAA NSLLIALICT QPALRNTSNF FLVSLFTSDL MVGLVVMPPA MLNALYGRWV LARGLCLLWT AFDVMCCSA SILNLCLISL DRYLLILSPL RYKLRMTPLR ALALVLGAWS LAALASFLPL LLGWHELGHA RPPVPGQCRL L ASLPFVLV ...String: GGSGWVAAAL CVVIALTAAA NSLLIALICT QPALRNTSNF FLVSLFTSDL MVGLVVMPPA MLNALYGRWV LARGLCLLWT AFDVMCCSA SILNLCLISL DRYLLILSPL RYKLRMTPLR ALALVLGAWS LAALASFLPL LLGWHELGHA RPPVPGQCRL L ASLPFVLV ASGLTFFLPS GAICFTYCRI LLAARKQAVQ VASLTTGMAS ADSRRLATKH SRKALKASLT LGILLGMFFV TW LPFFVAN IVQAVCDCIS PGLFDVLTWL GYCNSTMNPI IYPLFMRDFK RALGRFLPCP RCPRERQ UniProtKB: 5-hydroxytryptamine receptor 6 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.385957 KDa |
| Recombinant expression | Organism:  Insecta environmental sample (insect) Insecta environmental sample (insect) |
| Sequence | String: MHHHHHHENL YFQGGSSGSE LDQLRQEAEQ LKNQIRDARK ACADATLSQI TNNIDPVGRI QMRTRRTLRG HLAKIYAMHW GTDSRLLVS ASQDGKLIIW DSYTTNKVHA IPLRSSWVMT CAYAPSGNYV ACGGLDNICS IYNLKTREGN VRVSRELAGH T GYLSCCRF ...String: MHHHHHHENL YFQGGSSGSE LDQLRQEAEQ LKNQIRDARK ACADATLSQI TNNIDPVGRI QMRTRRTLRG HLAKIYAMHW GTDSRLLVS ASQDGKLIIW DSYTTNKVHA IPLRSSWVMT CAYAPSGNYV ACGGLDNICS IYNLKTREGN VRVSRELAGH T GYLSCCRF LDDNQIVTSS GDTTCALWDI ETGQQTTTFT GHTGDVMSLS LAPDTRLFVS GACDASAKLW DVREGMCRQT FT GHESDIN AICFFPNGNA FATGSDDATC RLFDLRADQE LMTYSHDNII CGITSVSFSK SGRLLLAGYD DFNCNVWDAL KAD RAGVLA GHDNRVSCLG VTDDGMAVAT GSWDSFLKIW N UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Insecta environmental sample (insect) Insecta environmental sample (insect) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 28.636793 KDa |
| Recombinant expression | Organism:  Insecta environmental sample (insect) Insecta environmental sample (insect) |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAAENLYFQ GHHHHHHHH |
-Macromolecule #5: Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit...
| Macromolecule | Name: Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.385246 KDa |
| Recombinant expression | Organism:  Insecta environmental sample (insect) Insecta environmental sample (insect) |
| Sequence | String: MGCTLSAEDK AAVERSKMIE KQLQKDKQVY RATHRLLLLG AGESGKSTIV KQMRILHVNG FNGDSEKATK VQDIKNNLKE AIETIVAAM SNLVPPVELA NPENQFRVDY ILSVMNVPDF DFPPEFYEHA KALWEDEGVR ACYERSNEYQ LIDCAQYFLD K IDVIKQAD ...String: MGCTLSAEDK AAVERSKMIE KQLQKDKQVY RATHRLLLLG AGESGKSTIV KQMRILHVNG FNGDSEKATK VQDIKNNLKE AIETIVAAM SNLVPPVELA NPENQFRVDY ILSVMNVPDF DFPPEFYEHA KALWEDEGVR ACYERSNEYQ LIDCAQYFLD K IDVIKQAD YVPSDQDLLR CRVLTSGIFE TKFQVDKVNF HMFDVGGQRD ERRKWIQCFN DVTAIIFVVA SSSYNMVIRE DN QTNRLQE ALNLFKSIWN NRWLRTISVI LFLNKQDLLA EKVLAGKSKI EDYFPEFARY TTPEDATPEP GEDPRVTRAK YFI RDEFLR ISTASGDGRH YCYPHFTCAV DTENIRRVFN DCRDIIQRMH LRQYELL UniProtKB: Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit alpha isoforms short |
-Macromolecule #6: SEROTONIN
| Macromolecule | Name: SEROTONIN / type: ligand / ID: 6 / Number of copies: 1 / Formula: SRO |
|---|---|
| Molecular weight | Theoretical: 176.215 Da |
| Chemical component information | 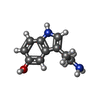 ChemComp-SRO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 388504 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller
























 Z
Z Y
Y X
X