[English] 日本語
 Yorodumi
Yorodumi- EMDB-33565: Structure of SUR2B in complex with Mg-ATP and repaglinide in the ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of SUR2B in complex with Mg-ATP and repaglinide in the inward-facing conformation | ||||||||||||
 Map data Map data | sharpened map of SUR2B in complex with Mg-ATP and RPG | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords |  SUR2B / SUR2B /  ABC transporter / ABC transporter /  repaglinide / repaglinide /  MEMBRANE PROTEIN MEMBRANE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsubstrate-dependent cell migration, cell contraction /  ATP sensitive Potassium channels / ABC-family proteins mediated transport / ATP sensitive Potassium channels / ABC-family proteins mediated transport /  circulatory system development / inward rectifying potassium channel / circulatory system development / inward rectifying potassium channel /  sulfonylurea receptor activity / cardiac conduction / ATPase-coupled monoatomic cation transmembrane transporter activity / coronary vasculature development / cardiac muscle cell contraction ...substrate-dependent cell migration, cell contraction / sulfonylurea receptor activity / cardiac conduction / ATPase-coupled monoatomic cation transmembrane transporter activity / coronary vasculature development / cardiac muscle cell contraction ...substrate-dependent cell migration, cell contraction /  ATP sensitive Potassium channels / ABC-family proteins mediated transport / ATP sensitive Potassium channels / ABC-family proteins mediated transport /  circulatory system development / inward rectifying potassium channel / circulatory system development / inward rectifying potassium channel /  sulfonylurea receptor activity / cardiac conduction / ATPase-coupled monoatomic cation transmembrane transporter activity / coronary vasculature development / cardiac muscle cell contraction / inorganic cation transmembrane transport / sulfonylurea receptor activity / cardiac conduction / ATPase-coupled monoatomic cation transmembrane transporter activity / coronary vasculature development / cardiac muscle cell contraction / inorganic cation transmembrane transport /  syntaxin binding / syntaxin binding /  action potential / Ion homeostasis / response to ATP / action potential / Ion homeostasis / response to ATP /  heterocyclic compound binding / potassium ion import across plasma membrane / blood vessel development / monoatomic cation transmembrane transport / ATPase-coupled transmembrane transporter activity / potassium channel regulator activity / ABC-type transporter activity / heart morphogenesis / potassium ion transmembrane transport / heterocyclic compound binding / potassium ion import across plasma membrane / blood vessel development / monoatomic cation transmembrane transport / ATPase-coupled transmembrane transporter activity / potassium channel regulator activity / ABC-type transporter activity / heart morphogenesis / potassium ion transmembrane transport /  T-tubule / negative regulation of blood pressure / T-tubule / negative regulation of blood pressure /  sarcomere / acrosomal vesicle / transmembrane transport / potassium ion transport / sarcomere / acrosomal vesicle / transmembrane transport / potassium ion transport /  regulation of blood pressure / fibroblast proliferation / defense response to virus / transmembrane transporter binding / response to xenobiotic stimulus / regulation of blood pressure / fibroblast proliferation / defense response to virus / transmembrane transporter binding / response to xenobiotic stimulus /  ATP hydrolysis activity / ATP hydrolysis activity /  ATP binding / ATP binding /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | ||||||||||||
| Biological species |   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) | ||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.73 Å cryo EM / Resolution: 3.73 Å | ||||||||||||
 Authors Authors | Chen L / Ding D / Hou T | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: The inhibition mechanism of the SUR2A-containing K channel by a regulatory helix. Authors: Dian Ding / Tianyi Hou / Miao Wei / Jing-Xiang Wu / Lei Chen /  Abstract: K channels are metabolic sensors for intracellular ATP/ADP ratios, play essential roles in many physiological processes, and are implicated in a spectrum of pathological conditions. SUR2A-containing ...K channels are metabolic sensors for intracellular ATP/ADP ratios, play essential roles in many physiological processes, and are implicated in a spectrum of pathological conditions. SUR2A-containing K channels differ from other subtypes in their sensitivity to Mg-ADP activation. However, the underlying structural mechanism remains poorly understood. Here we present a series of cryo-EM structures of SUR2A in the presence of different combinations of Mg-nucleotides and the allosteric inhibitor repaglinide. These structures uncover regulatory helix (R helix) on the NBD1-TMD2 linker, which wedges between NBD1 and NBD2. R helix stabilizes SUR2A in the NBD-separated conformation to inhibit channel activation. The competitive binding of Mg-ADP with Mg-ATP to NBD2 mobilizes the R helix to relieve such inhibition, allowing channel activation. The structures of SUR2B in similar conditions suggest that the C-terminal 42 residues of SUR2B enhance the structural dynamics of NBD2 and facilitate the dissociation of the R helix and the binding of Mg-ADP to NBD2, promoting NBD dimerization and subsequent channel activation. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33565.map.gz emd_33565.map.gz | 117.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33565-v30.xml emd-33565-v30.xml emd-33565.xml emd-33565.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33565.png emd_33565.png | 38.7 KB | ||
| Filedesc metadata |  emd-33565.cif.gz emd-33565.cif.gz | 6.5 KB | ||
| Others |  emd_33565_half_map_1.map.gz emd_33565_half_map_1.map.gz emd_33565_half_map_2.map.gz emd_33565_half_map_2.map.gz | 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33565 http://ftp.pdbj.org/pub/emdb/structures/EMD-33565 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33565 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33565 | HTTPS FTP |
-Related structure data
| Related structure data |  7y1lMC  7y1jC  7y1kC  7y1mC  7y1nC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33565.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33565.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map of SUR2B in complex with Mg-ATP and RPG | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.834 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half-A map of SUR2B in complex with Mg-ATP and RPG
| File | emd_33565_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-A map of SUR2B in complex with Mg-ATP and RPG | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-B map of SUR2B in complex with Mg-ATP and RPG
| File | emd_33565_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-B map of SUR2B in complex with Mg-ATP and RPG | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ATP-binding cassette sub-family C member 9, isoform B
| Entire | Name: ATP-binding cassette sub-family C member 9, isoform B |
|---|---|
| Components |
|
-Supramolecule #1: ATP-binding cassette sub-family C member 9, isoform B
| Supramolecule | Name: ATP-binding cassette sub-family C member 9, isoform B / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Molecular weight | Theoretical: 174 kDa/nm |
-Macromolecule #1: Isoform SUR2B of ATP-binding cassette sub-family C member 9
| Macromolecule | Name: Isoform SUR2B of ATP-binding cassette sub-family C member 9 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Molecular weight | Theoretical: 174.488562 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSLSFCGNNI SSYNIYHGVL QNPCFVDALN LVPHVFLLFI TFPILFIGWG SQSSKVQIHH NTWLHFPGHN LRWILTFALL FVHVCEIAE GIVSDSQRAS RHLHLFMPAV MGFVATTTSI VYYHNIETSN FPKLLLALFL YWVMAFITKT IKLVKYWQLG W GMSDLRFC ...String: MSLSFCGNNI SSYNIYHGVL QNPCFVDALN LVPHVFLLFI TFPILFIGWG SQSSKVQIHH NTWLHFPGHN LRWILTFALL FVHVCEIAE GIVSDSQRAS RHLHLFMPAV MGFVATTTSI VYYHNIETSN FPKLLLALFL YWVMAFITKT IKLVKYWQLG W GMSDLRFC ITGVMVILNG LLMAVEINVI RVRRYVFFMN PQKVKPPEDL QDLGVRFLQP FVNLLSKATY WWMNTLIISA HR KPIDLKA IGKLPIAMRA VTNYVCLKEA YEEQKKKAAD HPNRTPSIWL AMYRAFGRPI LLSSTFRYLA DLLGFAGPLC ISG IVQRVN EPKNNTTRFS ETLSSKEFLE NAHVLAVLLF LALILQRTFL QASYYVTIET GINLRGALLA MIYNKILRLS TSNL SMGEM TLGQINNLVA IETNQLMWFL FLCPNLWAMP VQIIMGVILL YNLLGSSALV GAAVIVLLAP IQYFIATKLA EAQKS TLDY STERLKKTNE ILKGIKLLKL YAWEHIFCKS VEETRMKELS SLKTFALYTS LSIFMNAAIP IAAVLATFVT HAYASG NNL KPAEAFASLS LFHILVTPLF LLSTVVRFAV KAIISVQKLN EFLLSDEIGE DSWRTGEGTL PFESCKKHTG VQSKPIN RK QPGRYHLDNY EQARRLRPAE TEDVAIKVTN GYFSWGSGLA TLSNIDIRIP TGQLTMIVGQ VGCGKSSLLL AILGEMQT L EGKVYWNNVN ESEPSFEATR SRSRYSVAYA AQKPWLLNAT VEENITFGSS FNRQRYKAVT DACSLQPDID LLPFGDQTE IGERGINLSG GQRQRICVAR ALYQNTNIVF LDDPFSALDI HLSDHLMQEG ILKFLQDDKR TVVLVTHKLQ YLTHADWIIA MKDGSVLRE GTLKDIQTKD VELYEHWKTL MNRQDQELEK DMEADQTTLE RKTLRRAMYS REAKAQMEDE DEEEEEEEDE D DNMSTVMR LRTKMPWKTC WWYLTSGGFF LLFLMIFSKL LKHSVIVAID YWLATWTSEY SINDPGKADQ TFYVAGFSIL CG AGIFLCL VTSLTVEWMG LTAAKNLHHN LLNKIILGPI RFFDTTPLGL ILNRFSADTN IIDQHIPPTL ESLTRSTLLC LSA IGMISY ATPVFLIALA PLGVAFYFIQ KYFRVASKDL QELDDSTQLP LLCHFSETAE GLTTIRAFRH ETRFKQRMLE LTDT NNIAY LFLSAANRWL EVRTDYLGAC IVLTASIASI SGSSNSGLVG LGLLYALTIT NYLNWVVRNL ADLEVQMGAV KKVNS FLTM ESENYEGTMD PSQVPEHWPQ EGEIKIHDLC VRYENNLKPV LKHVKAYIKP GQKVGICGRT GSGKSSLSLA FFRMVD IFD GKIVIDGIDI SKLPLHTLRS RLSIILQDPI LFSGSIRFNL DPECKCTDDR LWEALEIAQL KNMVKSLPGG LDATVTE GG ENFSVGQRQL FCLARAFVRK SSILIMDEAT ASIDMATENI LQKVVMTAFA DRTVVTIAHR VHTILTADLV IVMKRGNI L EYDTPESLLA QEDGVFASFV RADM UniProtKB: ATP-binding cassette sub-family C member 9 |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 2 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #4: Repaglinide
| Macromolecule | Name: Repaglinide / type: ligand / ID: 4 / Number of copies: 1 / Formula: BJX |
|---|---|
| Molecular weight | Theoretical: 452.586 Da |
| Chemical component information | 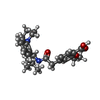 ChemComp-BJX: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.8 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.8 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 52.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.73 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 60550 |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X