+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of MrgD-Gi complex with beta-alanine (local) | |||||||||
 Map data Map data | post-process map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Complex / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationG protein-coupled receptor activity / electron transport chain / periplasmic space / electron transfer activity / iron ion binding / G protein-coupled receptor signaling pathway / heme binding / extracellular space / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Suzuki S / Iida M / Kawamoto A / Oshima A | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2022 Journal: Commun Biol / Year: 2022Title: Structural insight into the activation mechanism of MrgD with heterotrimeric Gi-protein revealed by cryo-EM. Authors: Shota Suzuki / Momoko Iida / Yoko Hiroaki / Kotaro Tanaka / Akihiro Kawamoto / Takayuki Kato / Atsunori Oshima /  Abstract: MrgD, a member of the Mas-related G protein-coupled receptor (MRGPR) family, has high basal activity for Gi activation. It recognizes endogenous ligands, such as β-alanine, and is involved in pain ...MrgD, a member of the Mas-related G protein-coupled receptor (MRGPR) family, has high basal activity for Gi activation. It recognizes endogenous ligands, such as β-alanine, and is involved in pain and itch signaling. The lack of a high-resolution structure for MrgD hinders our understanding of whether its activation is ligand-dependent or constitutive. Here, we report two cryo-EM structures of the MrgD-Gi complex in the β-alanine-bound and apo states at 3.1 Å and 2.8 Å resolution, respectively. These structures show that β-alanine is bound to a shallow pocket at the extracellular domains. The extracellular half of the sixth transmembrane helix undergoes a significant movement and is tightly packed into the third transmembrane helix through hydrophobic residues, creating the active form. Our structures demonstrate a structural basis for the characteristic ligand recognition of MrgD. These findings provide a framework to guide drug designs targeting the MrgD receptor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33556.map.gz emd_33556.map.gz | 117.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33556-v30.xml emd-33556-v30.xml emd-33556.xml emd-33556.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33556_fsc.xml emd_33556_fsc.xml | 14.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_33556.png emd_33556.png | 67 KB | ||
| Masks |  emd_33556_msk_1.map emd_33556_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-33556.cif.gz emd-33556.cif.gz | 6.1 KB | ||
| Others |  emd_33556_half_map_1.map.gz emd_33556_half_map_1.map.gz emd_33556_half_map_2.map.gz emd_33556_half_map_2.map.gz | 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33556 http://ftp.pdbj.org/pub/emdb/structures/EMD-33556 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33556 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33556 | HTTPS FTP |
-Validation report
| Summary document |  emd_33556_validation.pdf.gz emd_33556_validation.pdf.gz | 1004.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33556_full_validation.pdf.gz emd_33556_full_validation.pdf.gz | 1004.4 KB | Display | |
| Data in XML |  emd_33556_validation.xml.gz emd_33556_validation.xml.gz | 16.2 KB | Display | |
| Data in CIF |  emd_33556_validation.cif.gz emd_33556_validation.cif.gz | 21.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33556 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33556 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33556 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33556 | HTTPS FTP |
-Related structure data
| Related structure data |  7y14MC  7y12C  7y13C  7y15C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-11073 (Title: Cryo-EM structure of MrgD-Gi complex with beta-alanine EMPIAR-11073 (Title: Cryo-EM structure of MrgD-Gi complex with beta-alanineData size: 4.1 TB Data #1: K3 movies for MrgD-Gi complex with beta-alanine [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33556.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33556.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post-process map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.675 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_33556_msk_1.map emd_33556_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_33556_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_33556_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : beta-alanine bound MrgD-Gi complex with the scFv16
| Entire | Name: beta-alanine bound MrgD-Gi complex with the scFv16 |
|---|---|
| Components |
|
-Supramolecule #1: beta-alanine bound MrgD-Gi complex with the scFv16
| Supramolecule | Name: beta-alanine bound MrgD-Gi complex with the scFv16 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Soluble cytochrome b562,Mas-related G-protein coupled receptor me...
| Macromolecule | Name: Soluble cytochrome b562,Mas-related G-protein coupled receptor member D type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.141285 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDKADLEDN WETLNDNLKV IEKADNAAQV KDALTKMRAA ALDAQKATPP KLEDKSPDSP EMKDFRHGF DILVGQIDDA LKLANEGKVK EAQAAAEQLK TTRNAYIQKY LNSSGTVESA LNYSRGSTVH TAYLVLSSLA M FTCLCGMA ...String: MKTIIALSYI FCLVFADYKD DDDKADLEDN WETLNDNLKV IEKADNAAQV KDALTKMRAA ALDAQKATPP KLEDKSPDSP EMKDFRHGF DILVGQIDDA LKLANEGKVK EAQAAAEQLK TTRNAYIQKY LNSSGTVESA LNYSRGSTVH TAYLVLSSLA M FTCLCGMA GNSMVIWLLG FRMHRNPFCI YILNLAAADL LFLFSMASTL SLETQPLVNT TDKVHELMKR LMYFAYTVGL SL LTAISTQ RCLSVLFPIW FKCHRPRHLS AWVCGLLWTL CLLMNGLTSS FCSKFLKFNE DRCFRVDMVQ AALIMGVLTP VMT LSSLTL FVWVRRSSQQ WRRQPTRLFV VVLASVLVFL ICSLPLSIYW FVLYWLSLPP EMQVLCFSLS RLSSSVSSSA NPVI YFLVG SRRSHRLPTR SLGTVLQQAL REEPELEGGE TPTVGTNEMG A UniProtKB: Soluble cytochrome b562, Mas-related G-protein coupled receptor member D |
-Macromolecule #2: BETA-ALANINE
| Macromolecule | Name: BETA-ALANINE / type: ligand / ID: 2 / Number of copies: 1 / Formula: BAL |
|---|---|
| Molecular weight | Theoretical: 89.093 Da |
| Chemical component information | 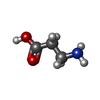 ChemComp-BAL: |
-Macromolecule #3: PALMITIC ACID
| Macromolecule | Name: PALMITIC ACID / type: ligand / ID: 3 / Number of copies: 4 / Formula: PLM |
|---|---|
| Molecular weight | Theoretical: 256.424 Da |
| Chemical component information |  ChemComp-PLM: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot time 3 seconds blot force 5. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: The Microscope implicated Cs corrector. |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.7000000000000001 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































