[English] 日本語
 Yorodumi
Yorodumi- EMDB-32588: The Cryo-EM structure of siphonaxanthin chlorophyll a/b type ligh... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
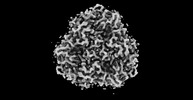
| Title | The Cryo-EM structure of siphonaxanthin chlorophyll a/b type light-harvesting complex II | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | light-harvesting complex II / siphonaxanthin / Chlorophyll b replacement / blue-green utilization / PHOTOSYNTHESIS | |||||||||
| Biological species |  Codium fragile (dead man's fingers) Codium fragile (dead man's fingers) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Seki S / Nakaniwa T / Castro-Hartmann P / Sader K / Kawamoto A / Tanaka H / Qian P / Kurisu G / Fujii R | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: BBA Adv / Year: 2022 Journal: BBA Adv / Year: 2022Title: Structural insights into blue-green light utilization by marine green algal light harvesting complex II at 2.78 Å. Authors: Soichiro Seki / Tetsuko Nakaniwa / Pablo Castro-Hartmann / Kasim Sader / Akihiro Kawamoto / Hideaki Tanaka / Pu Qian / Genji Kurisu / Ritsuko Fujii /   Abstract: Light-harvesting complex II (LHCII) present in plants and green algae absorbs solar energy to promote photochemical reactions. A marine green macroalga, , exhibits the unique characteristic of ...Light-harvesting complex II (LHCII) present in plants and green algae absorbs solar energy to promote photochemical reactions. A marine green macroalga, , exhibits the unique characteristic of absorbing blue-green light from the sun during photochemical reactions while being underwater owing to the presence of pigment-altered LHCII called siphonaxanthin-chlorophyll binding protein (SCP). In this study, we determined the structure of SCP at a resolution of 2.78 Å using cryogenic electron microscopy. SCP has a trimeric structure, wherein each monomer containing two lutein and two chlorophyll molecules in the plant-type LHCII are replaced by siphonaxanthin and its ester and two chlorophyll molecules, respectively. Siphonaxanthin occupies the binding site in SCP having a polarity in the trimeric inner core, and exhibits a distorted conjugated chain comprising a carbonyl group hydrogen bonded to a cysteine residue of apoprotein. These features suggest that the siphonaxanthin molecule is responsible for the characteristic green absorption of SCP. The replaced chlorophyll molecules extend the region of the stromal side chlorophyll cluster, spanning two adjacent monomers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32588.map.gz emd_32588.map.gz | 18.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32588-v30.xml emd-32588-v30.xml emd-32588.xml emd-32588.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32588_fsc.xml emd_32588_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_32588.png emd_32588.png | 42.4 KB | ||
| Filedesc metadata |  emd-32588.cif.gz emd-32588.cif.gz | 6.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32588 http://ftp.pdbj.org/pub/emdb/structures/EMD-32588 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32588 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32588 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_32588.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32588.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.662 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : siphonaxanthin chlorophyll a/b-type light-harvesting complex II
| Entire | Name: siphonaxanthin chlorophyll a/b-type light-harvesting complex II |
|---|---|
| Components |
|
-Supramolecule #1: siphonaxanthin chlorophyll a/b-type light-harvesting complex II
| Supramolecule | Name: siphonaxanthin chlorophyll a/b-type light-harvesting complex II type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Codium fragile (dead man's fingers) Codium fragile (dead man's fingers) |
| Molecular weight | Theoretical: 119 KDa |
-Macromolecule #1: siphonaxanthin chlorophyll a/b binding light-harvesting complex II
| Macromolecule | Name: siphonaxanthin chlorophyll a/b binding light-harvesting complex II type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Codium fragile (dead man's fingers) / Strain: KU-0654 Codium fragile (dead man's fingers) / Strain: KU-0654 |
| Molecular weight | Theoretical: 24.190102 KDa |
| Sequence | String: IEFYGPDRAL WLGPYSEGAV PSYLTGEFPG DYGWDSAGLS ADPETFAANR ELELIHARWA MLGVVGCLTP EALEKYSGVE FGEATWFKA GSQIFAEGGI DYLGNPSLVH AQSILAIVWS QVVLMGLAEG YRVSGGPLGE ATDPLYPGEA FDPFGFADDP E TFSELKIK ...String: IEFYGPDRAL WLGPYSEGAV PSYLTGEFPG DYGWDSAGLS ADPETFAANR ELELIHARWA MLGVVGCLTP EALEKYSGVE FGEATWFKA GSQIFAEGGI DYLGNPSLVH AQSILAIVWS QVVLMGLAEG YRVSGGPLGE ATDPLYPGEA FDPFGFADDP E TFSELKIK EIKNGRLAMF AMFGFFVQAL QTGKGPVECW ASHIEDPVAN NGFVYATKFF GAQIF |
-Macromolecule #2: CHLOROPHYLL B
| Macromolecule | Name: CHLOROPHYLL B / type: ligand / ID: 2 / Number of copies: 24 / Formula: CHL |
|---|---|
| Molecular weight | Theoretical: 907.472 Da |
| Chemical component information |  ChemComp-CHL: |
-Macromolecule #3: CHLOROPHYLL A
| Macromolecule | Name: CHLOROPHYLL A / type: ligand / ID: 3 / Number of copies: 15 / Formula: CLA |
|---|---|
| Molecular weight | Theoretical: 893.489 Da |
| Chemical component information |  ChemComp-CLA: |
-Macromolecule #4: (1R,3R)-6-{(3E,5E,7E,9E,11E,13E,15E,17E)-18-[(1S,4R,6R)-4-HYDROXY...
| Macromolecule | Name: (1R,3R)-6-{(3E,5E,7E,9E,11E,13E,15E,17E)-18-[(1S,4R,6R)-4-HYDROXY-2,2,6-TRIMETHYL-7-OXABICYCLO[4.1.0]HEPT-1-YL]-3,7,12,16-TETRAMETHYLOCTADECA-1,3,5,7,9,11,13,15,17-NONAENYLIDENE}-1,5,5-TRIMETHYLCYCLOHEXANE-1,3-DIOL type: ligand / ID: 4 / Number of copies: 3 / Formula: NEX |
|---|---|
| Molecular weight | Theoretical: 600.87 Da |
| Chemical component information |  ChemComp-NEX: |
-Macromolecule #5: Siphonein
| Macromolecule | Name: Siphonein / type: ligand / ID: 5 / Number of copies: 3 / Formula: 0UR |
|---|---|
| Molecular weight | Theoretical: 781.157 Da |
| Chemical component information |  ChemComp-0UR: |
-Macromolecule #6: Siphonaxanthin
| Macromolecule | Name: Siphonaxanthin / type: ligand / ID: 6 / Number of copies: 3 / Formula: 0IE |
|---|---|
| Molecular weight | Theoretical: 600.87 Da |
| Chemical component information |  ChemComp-0IE: |
-Macromolecule #7: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
| Macromolecule | Name: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE / type: ligand / ID: 7 / Number of copies: 3 / Formula: LHG |
|---|---|
| Molecular weight | Theoretical: 722.97 Da |
| Chemical component information |  ChemComp-LHG: |
-Macromolecule #8: water
| Macromolecule | Name: water / type: ligand / ID: 8 / Number of copies: 54 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8.2 Component:
| |||||||||
| Grid | Material: COPPER | |||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 2 / Number real images: 8935 / Average electron dose: 50.75 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 120000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: B / Chain - Residue range: 14-231 / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-7wlm: |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















