+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31361 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Mumps virus nucleocapsid ring | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nucleocapcid / Mumps virus / NUCLEAR PROTEIN | |||||||||
| Function / homology | Paramyxovirus nucleocapsid protein / Paramyxovirus nucleocapsid protein / helical viral capsid / viral nucleocapsid / host cell cytoplasm / ribonucleoprotein complex / structural molecule activity / RNA binding / Nucleoprotein Function and homology information Function and homology information | |||||||||
| Biological species |   Mumps virus (strain Jeryl-Lynn) / Mumps virus (strain Jeryl-Lynn) /  | |||||||||
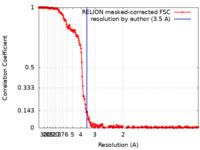
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Su X / Shen Q | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2021 Journal: Commun Biol / Year: 2021Title: Structural plasticity of mumps virus nucleocapsids with cryo-EM structures. Authors: Hong Shan / Xin Su / Tianhao Li / Yuqi Qin / Na Zhang / Liuyan Yang / Linsha Ma / Yun Bai / Lei Qi / Yunhui Liu / Qing-Tao Shen /  Abstract: Mumps virus (MuV) is a highly contagious human pathogen and frequently causes worldwide outbreaks despite available vaccines. Similar to other mononegaviruses such as Ebola and rabies, MuV uses a ...Mumps virus (MuV) is a highly contagious human pathogen and frequently causes worldwide outbreaks despite available vaccines. Similar to other mononegaviruses such as Ebola and rabies, MuV uses a single-stranded negative-sense RNA as its genome, which is enwrapped by viral nucleoproteins into the helical nucleocapsid. The nucleocapsid acts as a scaffold for genome condensation and as a template for RNA replication and transcription. Conformational changes in the MuV nucleocapsid are required to switch between different activities, but the underlying mechanism remains elusive due to the absence of high-resolution structures. Here, we report two MuV nucleoprotein-RNA rings with 13 and 14 protomers, one stacked-ring filament and two nucleocapsids with distinct helical pitches, in dense and hyperdense states, at near-atomic resolutions using cryo-electron microscopy. Structural analysis of these in vitro assemblies indicates that the C-terminal tail of MuV nucleoprotein likely regulates the assembly of helical nucleocapsids, and the C-terminal arm may be relevant for the transition between the dense and hyperdense states of helical nucleocapsids. Our results provide the molecular mechanism for structural plasticity among different MuV nucleocapsids and create a possible link between structural plasticity and genome condensation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31361.map.gz emd_31361.map.gz | 845 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31361-v30.xml emd-31361-v30.xml emd-31361.xml emd-31361.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_31361_fsc.xml emd_31361_fsc.xml | 23.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_31361.png emd_31361.png | 134 KB | ||
| Filedesc metadata |  emd-31361.cif.gz emd-31361.cif.gz | 5.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31361 http://ftp.pdbj.org/pub/emdb/structures/EMD-31361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31361 | HTTPS FTP |
-Related structure data
| Related structure data |  7ewqMC  7exaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31361.map.gz / Format: CCP4 / Size: 1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31361.map.gz / Format: CCP4 / Size: 1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.53 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mumps virus nocleocapcid
| Entire | Name: Mumps virus nocleocapcid |
|---|---|
| Components |
|
-Supramolecule #1: Mumps virus nocleocapcid
| Supramolecule | Name: Mumps virus nocleocapcid / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Nucleoprotein
| Macromolecule | Name: Nucleoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mumps virus (strain Jeryl-Lynn) / Strain: Jeryl-Lynn Mumps virus (strain Jeryl-Lynn) / Strain: Jeryl-Lynn |
| Molecular weight | Theoretical: 61.470008 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSSVLKAFER FTIEQELQDR GEEGSIPPET LKSAVKVFVI NTPNPTTRYQ MLNFCLRIIC SQNARASHRV GALITLFSLP SAGMQNHIR LADRSPEAQI ERCEIDGFEP GTYRLIPNAR ANLTANEIAA YALLADDLPP TINNGTPYVH ADVEGQPCDE I EQFLDRCY ...String: MSSVLKAFER FTIEQELQDR GEEGSIPPET LKSAVKVFVI NTPNPTTRYQ MLNFCLRIIC SQNARASHRV GALITLFSLP SAGMQNHIR LADRSPEAQI ERCEIDGFEP GTYRLIPNAR ANLTANEIAA YALLADDLPP TINNGTPYVH ADVEGQPCDE I EQFLDRCY SVLIQAWVMV CKCMTAYDQP AGSADRRFAK YQQQGRLEAR YMLQPEAQRL IQTAIRKSLV VRQYLTFELQ LA RRQGLLS NRYYAMVGDI GKYIENSGLT AFFLTLKYAL GTKWSPLSLA AFTGELTKLR SLMMLYRGLG EQARYLALLE APQ IMDFAP GGYPLIFSYA MGVGTVLDVQ MRNYTYARPF LNGYYFQIGV ETARRQQGTV DNRVADDLGL TPEQRTEVTQ LVDR LARGR GAGIPGGPVN PFVPPVQQQQ PAAVYEDIPA LEESDDDGDE DGGAGFQNGV QLPAVRQGGQ TDFRAQPLQD PIQAQ LFMP LYPQVSNMPN NQNHQINRIG GLEHQDLLRY NENGDSQQDA RGEHVNTFPN NPNQNAQLQV GDWDE UniProtKB: Nucleoprotein |
-Macromolecule #2: RNA (5'-R(P*UP*UP*UP*UP*UP*U)-3')
| Macromolecule | Name: RNA (5'-R(P*UP*UP*UP*UP*UP*U)-3') / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.792037 KDa |
| Sequence | String: UUUUUU |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)