[English] 日本語
 Yorodumi
Yorodumi- EMDB-29484: E. coli 70S ribosome with an improved MS2 tag inserted in H98 (50... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli 70S ribosome with an improved MS2 tag inserted in H98 (50S focused map) | |||||||||
 Map data Map data | E. coli 70S ribosome with an improved MS2 tag inserted in H98 (50S focused map) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MS2-tag / H98 / ribosome | |||||||||
| Biological species |  | |||||||||
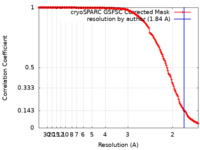
| Method | single particle reconstruction / cryo EM / Resolution: 1.84 Å | |||||||||
 Authors Authors | Nissley AJ / Cate JHD | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: RNA / Year: 2023 Journal: RNA / Year: 2023Title: Interactions between terminal ribosomal RNA helices stabilize the large ribosomal subunit. Authors: Amos J Nissley / Tammam S Kamal / Jamie H D Cate /  Abstract: The ribosome is a large ribonucleoprotein assembly that uses diverse and complex molecular interactions to maintain proper folding. In vivo assembled ribosomes have been isolated using MS2 tags ...The ribosome is a large ribonucleoprotein assembly that uses diverse and complex molecular interactions to maintain proper folding. In vivo assembled ribosomes have been isolated using MS2 tags installed in either the 16S or 23S ribosomal RNAs (rRNAs), to enable studies of ribosome structure and function in vitro. RNA tags in the 50S subunit have commonly been inserted into an extended helix H98 in 23S rRNA, as this addition does not affect cellular growth or in vitro ribosome activity. Here, we find that 50S subunits with MS2 tags inserted in H98 are destabilized compared to wild-type (WT) 50S subunits. We identify the loss of RNA-RNA tertiary contacts that bridge helices H1, H94, and H98 as the cause of destabilization. Using cryogenic electron microscopy (cryo-EM), we show that this interaction is disrupted by the addition of the MS2 tag and can be restored through the insertion of a single adenosine in the extended H98 helix. This work establishes ways to improve MS2 tags in the 50S subunit that maintain ribosome stability and investigates a complex RNA tertiary structure that may be important for stability in various bacterial ribosomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29484.map.gz emd_29484.map.gz | 324.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29484-v30.xml emd-29484-v30.xml emd-29484.xml emd-29484.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29484_fsc.xml emd_29484_fsc.xml | 14.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_29484.png emd_29484.png | 122.9 KB | ||
| Filedesc metadata |  emd-29484.cif.gz emd-29484.cif.gz | 4.1 KB | ||
| Others |  emd_29484_half_map_1.map.gz emd_29484_half_map_1.map.gz emd_29484_half_map_2.map.gz emd_29484_half_map_2.map.gz | 318.1 MB 318.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29484 http://ftp.pdbj.org/pub/emdb/structures/EMD-29484 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29484 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29484 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29484.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29484.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli 70S ribosome with an improved MS2 tag inserted in H98 (50S focused map) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8248 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: E. coli 70S ribosome with an improved MS2...
| File | emd_29484_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli 70S ribosome with an improved MS2 tag inserted in H98 (50S focused half map) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: E. coli 70S ribosome with an improved MS2...
| File | emd_29484_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli 70S ribosome with an improved MS2 tag inserted in H98 (50S focused half map) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : E. coli 70S ribosome
| Entire | Name: E. coli 70S ribosome |
|---|---|
| Components |
|
-Supramolecule #1: E. coli 70S ribosome
| Supramolecule | Name: E. coli 70S ribosome / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: 50S Subunit
| Supramolecule | Name: 50S Subunit / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: 30S Subunit
| Supramolecule | Name: 30S Subunit / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.27 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 11736 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)