+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Human TRPM2 ion channel in 1 mM dADPR | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | TRPM2 / TRP / ADPR / ADP-ribose / NUDT9H / NUDT9 / calcium / ion channel / MEMBRANE PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報mono-ADP-D-ribose binding / manganese ion transmembrane transporter activity / dendritic cell differentiation / zinc ion transmembrane transport / response to purine-containing compound / cellular response to temperature stimulus / regulation of filopodium assembly / ligand-gated calcium channel activity / response to hydroperoxide / dendritic cell chemotaxis ...mono-ADP-D-ribose binding / manganese ion transmembrane transporter activity / dendritic cell differentiation / zinc ion transmembrane transport / response to purine-containing compound / cellular response to temperature stimulus / regulation of filopodium assembly / ligand-gated calcium channel activity / response to hydroperoxide / dendritic cell chemotaxis / TRP channels / calcium ion transmembrane import into cytosol / sodium channel activity / temperature homeostasis / intracellularly gated calcium channel activity / tertiary granule membrane / calcium ion import across plasma membrane / ficolin-1-rich granule membrane / specific granule membrane / monoatomic cation channel activity / cell projection / release of sequestered calcium ion into cytosol / cellular response to calcium ion / cytoplasmic vesicle membrane / regulation of actin cytoskeleton organization / calcium ion transmembrane transport / calcium channel activity / cellular response to hydrogen peroxide / calcium ion transport / response to heat / protein homotetramerization / perikaryon / lysosome / lysosomal membrane / calcium ion binding / Neutrophil degranulation / plasma membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 5.6 Å | |||||||||
 データ登録者 データ登録者 | Wang L / Fu TM / Xia S / Wu H | |||||||||
| 資金援助 | 1件
| |||||||||
 引用 引用 |  ジャーナル: To Be Published ジャーナル: To Be Publishedタイトル: A unified mechanism for human TRPM2 activation, desensitization and inhibition 著者: Wang L / Fu T-M | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_27923.map.gz emd_27923.map.gz | 94.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-27923-v30.xml emd-27923-v30.xml emd-27923.xml emd-27923.xml | 17.7 KB 17.7 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_27923.png emd_27923.png | 115.8 KB | ||
| Filedesc metadata |  emd-27923.cif.gz emd-27923.cif.gz | 6.6 KB | ||
| その他 |  emd_27923_half_map_1.map.gz emd_27923_half_map_1.map.gz emd_27923_half_map_2.map.gz emd_27923_half_map_2.map.gz | 76.1 MB 76 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27923 http://ftp.pdbj.org/pub/emdb/structures/EMD-27923 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27923 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27923 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_27923.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_27923.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #1
| ファイル | emd_27923_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_27923_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Human TRPM2 in 1 mM ADPR
| 全体 | 名称: Human TRPM2 in 1 mM ADPR |
|---|---|
| 要素 |
|
-超分子 #1: Human TRPM2 in 1 mM ADPR
| 超分子 | 名称: Human TRPM2 in 1 mM ADPR / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: Transient receptor potential cation channel subfamily M member 2
| 分子 | 名称: Transient receptor potential cation channel subfamily M member 2 タイプ: protein_or_peptide / ID: 1 / コピー数: 4 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 171.416188 KDa |
| 組換発現 | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 配列 | 文字列: MEPSALRKAG SEQEEGFEGL PRRVTDLGMV SNLRRSNSSL FKSWRLQCPF GNNDKQESLS SWIPENIKKK ECVYFVESSK LSDAGKVVC QCGYTHEQHL EEATKPHTFQ GTQWDPKKHV QEMPTDAFGD IVFTGLSQKV KKYVRVSQDT PSSVIYHLMT Q HWGLDVPN ...文字列: MEPSALRKAG SEQEEGFEGL PRRVTDLGMV SNLRRSNSSL FKSWRLQCPF GNNDKQESLS SWIPENIKKK ECVYFVESSK LSDAGKVVC QCGYTHEQHL EEATKPHTFQ GTQWDPKKHV QEMPTDAFGD IVFTGLSQKV KKYVRVSQDT PSSVIYHLMT Q HWGLDVPN LLISVTGGAK NFNMKPRLKS IFRRGLVKVA QTTGAWIITG GSHTGVMKQV GEAVRDFSLS SSYKEGELIT IG VATWGTV HRREGLIHPT GSFPAEYILD EDGQGNLTCL DSNHSHFILV DDGTHGQYGV EIPLRTRLEK FISEQTKERG GVA IKIPIV CVVLEGGPGT LHTIDNATTN GTPCVVVEGS GRVADVIAQV ANLPVSDITI SLIQQKLSVF FQEMFETFTE SRIV EWTKK IQDIVRRRQL LTVFREGKDG QQDVDVAILQ ALLKASRSQD HFGHENWDHQ LKLAVAWNRV DIARSEIFMD EWQWK PSDL HPTMTAALIS NKPEFVKLFL ENGVQLKEFV TWDTLLYLYE NLDPSCLFHS KLQKVLVEDP ERPACAPAAP RLQMHH VAQ VLRELLGDFT QPLYPRPRHN DRLRLLLPVP HVKLNVQGVS LRSLYKRSSG HVTFTMDPIR DLLIWAIVQN RRELAGI IW AQSQDCIAAA LACSKILKEL SKEEEDTDSS EEMLALAEEY EHRAIGVFTE CYRKDEERAQ KLLTRVSEAW GKTTCLQL A LEAKDMKFVS HGGIQAFLTK VWWGQLSVDN GLWRVTLCML AFPLLLTGLI SFREKRLQDV GTPAARARAF FTAPVVVFH LNILSYFAFL CLFAYVLMVD FQPVPSWCEC AIYLWLFSLV CEEMRQLFYD PDECGLMKKA ALYFSDFWNK LDVGAILLFV AGLTCRLIP ATLYPGRVIL SLDFILFCLR LMHIFTISKT LGPKIIIVKR MMKDVFFFLF LLAVWVVSFG VAKQAILIHN E RRVDWLFR GAVYHSYLTI FGQIPGYIDG VNFNPEHCSP NGTDPYKPKC PESDATQQRP AFPEWLTVLL LCLYLLFTNI LL LNLLIAM FNYTFQQVQE HTDQIWKFQR HDLIEEYHGR PAAPPPFILL SHLQLFIKRV VLKTPAKRHK QLKNKLEKNE EAA LLSWEI YLKENYLQNR QFQQKQRPEQ KIEDISNKVD AMVDLLDLDP LKRSGSMEQR LASLEEQVAQ TAQALHWIVR TLRA SGFSS EADVPTLASQ KAAEEPDAEP GGRKKTEEPG DSYHVNARHL LYPNCPVTRF PVPNEKVPWE TEFLIYDPPF YTAER KDAA AMDPMGDTLE PLSTIQYNVV DGLRDRRSFH GPYTVQAGLP LNPMGRTGLR GRGSLSCFGP NHTLYPMVTR WRRNED GAI CRKSIKKMLE VLVVKLPLSE HWALPGGSRE PGEMLPRKLK RILRQEHWPS FENLLKCGME VYKGYMDDPR NTDNAWI ET VAVSVHFQDQ NDVELNRLNS NLHACDSGAS IRWQVVDRRI PLYANHKTLL QKAAAEFGAH Y UniProtKB: Transient receptor potential cation channel subfamily M member 2 |
-分子 #2: ADENOSINE-5-DIPHOSPHORIBOSE
| 分子 | 名称: ADENOSINE-5-DIPHOSPHORIBOSE / タイプ: ligand / ID: 2 / コピー数: 4 / 式: APR |
|---|---|
| 分子量 | 理論値: 559.316 Da |
| Chemical component information | 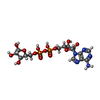 ChemComp-APR: |
-分子 #3: [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]ME...
| 分子 | 名称: [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL[HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE タイプ: ligand / ID: 3 / コピー数: 4 / 式: AR6 |
|---|---|
| 分子量 | 理論値: 559.316 Da |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.75 mg/mL |
|---|---|
| 緩衝液 | pH: 7.4 |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 42.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.2 µm / 最小 デフォーカス(公称値): 0.8 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)