[English] 日本語
 Yorodumi
Yorodumi- EMDB-27896: The closed C0-state flycatcher TRPM8 structure in complex with PI... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The closed C0-state flycatcher TRPM8 structure in complex with PI(4,5)P2 | |||||||||
 Map data Map data | Full map, sharpened with B-factor -50 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRPM8 / menthol receptor / cold receptor / cold sensor / PI(4 / 5)P2 / cooling agonists / temperature sensing / ion channel / sensory transduction / transient receptor potential ion channel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationligand-gated calcium channel activity / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Ficedula albicollis (Collared flycatcher) Ficedula albicollis (Collared flycatcher) | |||||||||
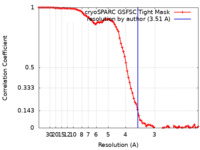
| Method | single particle reconstruction / cryo EM / Resolution: 3.51 Å | |||||||||
 Authors Authors | Yin Y / Zhang F / Feng S / Butay KJ / Borgnia MJ / Im W / Lee S-Y | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Activation mechanism of the mouse cold-sensing TRPM8 channel by cooling agonist and PIP. Authors: Ying Yin / Feng Zhang / Shasha Feng / Kevin John Butay / Mario J Borgnia / Wonpil Im / Seok-Yong Lee /  Abstract: The transient receptor potential melastatin 8 (TRPM8) channel is the primary molecular transducer responsible for the cool sensation elicited by menthol and cold in mammals. TRPM8 activation is ...The transient receptor potential melastatin 8 (TRPM8) channel is the primary molecular transducer responsible for the cool sensation elicited by menthol and cold in mammals. TRPM8 activation is controlled by cooling compounds together with the membrane lipid phosphatidylinositol 4,5-bisphosphate (PIP). Our knowledge of cold sensation and the therapeutic potential of TRPM8 for neuroinflammatory diseases and pain will be enhanced by understanding the structural basis of cooling agonist- and PIP-dependent TRPM8 activation. We present cryo-electron microscopy structures of mouse TRPM8 in closed, intermediate, and open states along the ligand- and PIP-dependent gating pathway. Our results uncover two discrete agonist sites, state-dependent rearrangements in the gate positions, and a disordered-to-ordered transition of the gate-forming S6-elucidating the molecular basis of chemically induced cool sensation in mammals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27896.map.gz emd_27896.map.gz | 32.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27896-v30.xml emd-27896-v30.xml emd-27896.xml emd-27896.xml | 19 KB 19 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27896_fsc.xml emd_27896_fsc.xml | 9.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_27896.png emd_27896.png | 27.3 KB | ||
| Masks |  emd_27896_msk_1.map emd_27896_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27896.cif.gz emd-27896.cif.gz | 6.8 KB | ||
| Others |  emd_27896_half_map_1.map.gz emd_27896_half_map_1.map.gz emd_27896_half_map_2.map.gz emd_27896_half_map_2.map.gz | 59 MB 59 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27896 http://ftp.pdbj.org/pub/emdb/structures/EMD-27896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27896 | HTTPS FTP |
-Related structure data
| Related structure data |  8e4qMC  8e4lC  8e4mC  8e4nC  8e4oC  8e4pC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27896.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27896.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map, sharpened with B-factor -50 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27896_msk_1.map emd_27896_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_27896_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_27896_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Transient receptor potential cation channel subfamily M member 8
| Entire | Name: Transient receptor potential cation channel subfamily M member 8 |
|---|---|
| Components |
|
-Supramolecule #1: Transient receptor potential cation channel subfamily M member 8
| Supramolecule | Name: Transient receptor potential cation channel subfamily M member 8 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Ficedula albicollis (Collared flycatcher) Ficedula albicollis (Collared flycatcher) |
-Macromolecule #1: Transient receptor potential cation channel subfamily M member 8
| Macromolecule | Name: Transient receptor potential cation channel subfamily M member 8 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ficedula albicollis (Collared flycatcher) Ficedula albicollis (Collared flycatcher) |
| Molecular weight | Theoretical: 131.343406 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MATLFQVSMG SMRHRRNGNF ESSRLLYSSM SRSIDVACSD ADLANFIQEN FKKRECVFFT KDTKSMGNLC KCGYPENQHI EGTQVNTTE KWNYKKHTKE LPTDAFGDIQ FENLGKRGKY IRLSCDTDSE TLYDLMTQHW HLKTPNLVIS VTGGAKNFAL K PRMRKIFS ...String: MATLFQVSMG SMRHRRNGNF ESSRLLYSSM SRSIDVACSD ADLANFIQEN FKKRECVFFT KDTKSMGNLC KCGYPENQHI EGTQVNTTE KWNYKKHTKE LPTDAFGDIQ FENLGKRGKY IRLSCDTDSE TLYDLMTQHW HLKTPNLVIS VTGGAKNFAL K PRMRKIFS RLIYIAQSKG AWIFTGGTHY GLMKYIGEVV RDNTISRSSE ENVVAIGIAA WGMISNRETL IRTADSDGSY LA HYIMDDL KRDPLYCLDN NHTHLLLVDN GTHGHPTIEA KVRTQLEKYI SERVIPESNY GGKIPIVCFA QGGGKETLKS INV AIKSKI PCVVVEGSGR IADVIASLVE AEGTLASSCV KESLLRFLPR TISRLSEEET ESWIKWIKEV LESPHLLTVI KIEE AGDEI VSNAISFALY KAFSTNEHDR DNWNGQLKLL LEWNQLDLAS DEIFTNDRNW ESADLQDVMF TALVKDRPKF VRLFL ENGL NLRKFLTTEV LRELYTNNFS SLVFKNLQIA KNSYNDALLT FVWKMVEDFR RGAKRDDKNS KDEMEIELSE ECPITR HPL QALFIWSVLQ NKKELSKVIW EQTRGCTLAA LGASKLLKSM AKVKNDINAA GESEELANEY ETRAVELFTE CYSNDED LA EQLLTYSCEA WGGSNCLELA VEARDQQFIA QPGVQNFLSK QWYGEISRDT KNWKIILCLF FFPLIGCGFI SFRKKPVE K TKKLFLYYVS FFTSPFVVFS WNVIFYIAFL LLFAYVLLMD FQKEPTALEI ILYVLVFILL CDEVRQWYMN GSKYFSDLW NVMDTLAIFY FIAGIVFRLH SDESSWYSGR VIFCLDYIVF TLRLIHIFTV SRNLGPKIIM LQRMMIDVFF FLFLFAVWMV AFGVARQGI LRKNEHRWEW IFRSVIYEPY LAMFGQYPDD IDGTTYNFDH CTFSGNESKP LCVELDANNQ PRFPEWITIP L VCIYMLST NILLVNLLVA MFGYTVGSVQ ENNDQVWKFQ RFFLVQEYCS RLTIPFPFVI FAYIFMVMRK CFKCCCKKES KE PSVCCSR NEDNEILAWE AVMKENYLVK INTKASDSSE EMVHRFRQLD AKLSDLKGLL KEISSKIKSN SLEVLFQGPD YKD DDDKAH HHHHHHHHH UniProtKB: Transient receptor potential cation channel subfamily M member 8 |
-Macromolecule #2: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(o...
| Macromolecule | Name: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phosphoryl]oxy-propyl] octanoate type: ligand / ID: 2 / Number of copies: 4 / Formula: PIO |
|---|---|
| Molecular weight | Theoretical: 746.566 Da |
| Chemical component information |  ChemComp-PIO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 4283 / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.25 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)