[English] 日本語
 Yorodumi
Yorodumi- EMDB-27875: Full-length APOBEC3G in complex with HIV-1 Vif, CBF-beta, and fork RNA -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full-length APOBEC3G in complex with HIV-1 Vif, CBF-beta, and fork RNA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Viral protein - human protein complex / ribonucleoprotein complex / VIRAL PROTEIN-RNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology information: / RUNX3 regulates RUNX1-mediated transcription / RUNX1 regulates transcription of genes involved in BCR signaling / cytidine deamination / RUNX1 regulates transcription of genes involved in interleukin signaling / RUNX2 regulates bone development / core-binding factor complex / RUNX1 regulates expression of components of tight junctions / positive regulation of CD8-positive, alpha-beta T cell differentiation / RUNX2 regulates chondrocyte maturation ...: / RUNX3 regulates RUNX1-mediated transcription / RUNX1 regulates transcription of genes involved in BCR signaling / cytidine deamination / RUNX1 regulates transcription of genes involved in interleukin signaling / RUNX2 regulates bone development / core-binding factor complex / RUNX1 regulates expression of components of tight junctions / positive regulation of CD8-positive, alpha-beta T cell differentiation / RUNX2 regulates chondrocyte maturation / single-stranded DNA cytosine deaminase / negative regulation of CD4-positive, alpha-beta T cell differentiation / DNA cytosine deamination / : / lymphocyte differentiation / cytidine to uridine editing / cytidine deaminase activity / RUNX1 and FOXP3 control the development of regulatory T lymphocytes (Tregs) / negative regulation of single stranded viral RNA replication via double stranded DNA intermediate / : / RUNX2 regulates genes involved in cell migration / RUNX2 regulates genes involved in differentiation of myeloid cells / Transcriptional regulation by RUNX2 / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / myeloid cell differentiation / RUNX3 Regulates Immune Response and Cell Migration / definitive hemopoiesis / RUNX1 regulates transcription of genes involved in differentiation of myeloid cells / Regulation of RUNX1 Expression and Activity / RUNX1 regulates transcription of genes involved in WNT signaling / RUNX1 regulates estrogen receptor mediated transcription / RUNX1 interacts with co-factors whose precise effect on RUNX1 targets is not known / RUNX2 regulates osteoblast differentiation / RUNX3 regulates p14-ARF / cell maturation / viral life cycle / virion component / Regulation of RUNX3 expression and activity / P-body / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Transcriptional regulation of granulopoiesis / osteoblast differentiation / protein polyubiquitination / Regulation of RUNX2 expression and activity / RUNX1 regulates transcription of genes involved in differentiation of HSCs / defense response to virus / Estrogen-dependent gene expression / sequence-specific DNA binding / host cell cytoplasm / transcription by RNA polymerase II / transcription coactivator activity / ribonucleoprotein complex / innate immune response / regulation of transcription by RNA polymerase II / host cell plasma membrane / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / RNA binding / zinc ion binding / nucleoplasm / membrane / nucleus / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /   Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
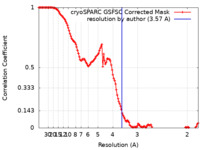
| Method | single particle reconstruction / cryo EM / Resolution: 3.57 Å | |||||||||
 Authors Authors | Ito F / Alvarez-Cabrera AL / Liu S / Yang H / Shiriaeva A / Zhou ZH / Chen XS | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Structural basis for HIV-1 antagonism of host APOBEC3G via Cullin E3 ligase. Authors: Fumiaki Ito / Ana L Alvarez-Cabrera / Shiheng Liu / Hanjing Yang / Anna Shiriaeva / Z Hong Zhou / Xiaojiang S Chen /  Abstract: Human APOBEC3G (A3G) is a virus restriction factor that inhibits HIV-1 replication and triggers lethal hypermutation on viral reverse transcripts. HIV-1 viral infectivity factor (Vif) breaches this ...Human APOBEC3G (A3G) is a virus restriction factor that inhibits HIV-1 replication and triggers lethal hypermutation on viral reverse transcripts. HIV-1 viral infectivity factor (Vif) breaches this host A3G immunity by hijacking a cellular E3 ubiquitin ligase complex to target A3G for ubiquitination and degradation. The molecular mechanism of A3G targeting by Vif-E3 ligase is unknown, limiting the antiviral efforts targeting this host-pathogen interaction crucial for HIV-1 infection. Here, we report the cryo-electron microscopy structures of A3G bound to HIV-1 Vif in complex with T cell transcription cofactor CBF-β and multiple components of the Cullin-5 RING E3 ubiquitin ligase. The structures reveal unexpected RNA-mediated interactions of Vif with A3G primarily through A3G's noncatalytic domain, while A3G's catalytic domain is poised for ubiquitin transfer. These structures elucidate the molecular mechanism by which HIV-1 Vif hijacks the host ubiquitin ligase to specifically target A3G to establish infection and offer structural information for the rational development of antiretroviral therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27875.map.gz emd_27875.map.gz | 108.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27875-v30.xml emd-27875-v30.xml emd-27875.xml emd-27875.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27875_fsc.xml emd_27875_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27875.png emd_27875.png | 69.1 KB | ||
| Filedesc metadata |  emd-27875.cif.gz emd-27875.cif.gz | 6.3 KB | ||
| Others |  emd_27875_half_map_1.map.gz emd_27875_half_map_1.map.gz emd_27875_half_map_2.map.gz emd_27875_half_map_2.map.gz | 200.4 MB 200.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27875 http://ftp.pdbj.org/pub/emdb/structures/EMD-27875 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27875 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27875 | HTTPS FTP |
-Validation report
| Summary document |  emd_27875_validation.pdf.gz emd_27875_validation.pdf.gz | 716.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27875_full_validation.pdf.gz emd_27875_full_validation.pdf.gz | 715.8 KB | Display | |
| Data in XML |  emd_27875_validation.xml.gz emd_27875_validation.xml.gz | 21.4 KB | Display | |
| Data in CIF |  emd_27875_validation.cif.gz emd_27875_validation.cif.gz | 27.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27875 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27875 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27875 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27875 | HTTPS FTP |
-Related structure data
| Related structure data |  8e40MC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27875.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27875.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_27875_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
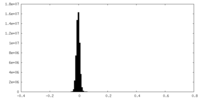
| Density Histograms |
-Half map: #1
| File | emd_27875_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of APOBEC3G with HIV-1 Vif and CBF-beta bound to fork RNA
| Entire | Name: Complex of APOBEC3G with HIV-1 Vif and CBF-beta bound to fork RNA |
|---|---|
| Components |
|
-Supramolecule #1: Complex of APOBEC3G with HIV-1 Vif and CBF-beta bound to fork RNA
| Supramolecule | Name: Complex of APOBEC3G with HIV-1 Vif and CBF-beta bound to fork RNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 99 KDa |
-Macromolecule #1: DNA dC->dU-editing enzyme APOBEC-3G
| Macromolecule | Name: DNA dC->dU-editing enzyme APOBEC-3G / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: single-stranded DNA cytosine deaminase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 45.18743 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGGSGGMKP QIRNMVEPMD PRTFVSNFNN RPILSGLDTV WLCCEVKTKD PSGPPLDAKI FQGKVYPKAK YHPEMRFLRW FHKWRQLHH DQEYKVTWYV SWSPCTRCAN SVATFLAKDP KVTLTIFVAR LYYFWDPDYQ QALRILAEAG ATMKIMNYNE F QDCWNKFV ...String: GPGGSGGMKP QIRNMVEPMD PRTFVSNFNN RPILSGLDTV WLCCEVKTKD PSGPPLDAKI FQGKVYPKAK YHPEMRFLRW FHKWRQLHH DQEYKVTWYV SWSPCTRCAN SVATFLAKDP KVTLTIFVAR LYYFWDPDYQ QALRILAEAG ATMKIMNYNE F QDCWNKFV DGRGKPFKPW NNLPKHYTLL QATLGELLRH LMDPGTFTSN FNNKPWVSGQ HETYLCYKVE RLHNDTWVPL NQ HRGFLRN QAPNIHGFPK GRHAQLCFLD LIPFWKLDGQ QYRVTCFTSW SPCFSCAQEM AKFISNNEHV SLCIFAARIY DDQ GRYQEG LRTLHRDGAK IAMMNYSEFE YCWDTFVDRQ GRPFQPWDGL DEHSQALSGR LRAILQNQGN UniProtKB: DNA dC->dU-editing enzyme APOBEC-3G |
-Macromolecule #2: Virion infectivity factor
| Macromolecule | Name: Virion infectivity factor / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 21.136408 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPMENRWQVM IVWQVDRMRI NTWKRLVKHH MYISRKAKDW FYRHHYESTN PKISSEVHIP LGDAKLVITT YWGLHTGERD WHLGQGVSI EWRKKRYSTQ VDPDLADQLI HLHYFDCFSE SAIRNTILGR IVSPRCEYQA GHNKVGSLQY LALAALIKPK Q IKPPLPSV RKLTEDRWNK UniProtKB: Virion infectivity factor |
-Macromolecule #3: Core-binding factor subunit beta
| Macromolecule | Name: Core-binding factor subunit beta / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.643814 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPRVVPDQRS KFENEEFFRK LSRECEIKYT GFRDRPHEER QARFQNACRD GRSEIAFVAT GTNLSLQFFP ASWQGEQRQT PSREYVDLE REAGKVYLKA PMILNGVCVI WKGWIDLQRL DGMGCLEFDE ERAQQEDALA QQAFEEARRR TREFEDRD UniProtKB: Core-binding factor subunit beta |
-Macromolecule #4: RNA
| Macromolecule | Name: RNA / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.875011 KDa |
| Sequence | String: UUUUUUUUUU UUUUAAAAAA AA |
-Macromolecule #5: RNA
| Macromolecule | Name: RNA / type: rna / ID: 5 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.00087 KDa |
| Sequence | String: UUUUUUUUAA AAAAAAAAAA AAAAA |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.15 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 11803 / Average exposure time: 8.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 150000 |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X