+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Citrus V-ATPase State 1, H in contact with subunits AB | |||||||||
 Map data Map data | Citrus V-ATPase State 1, H in contact with subunits AB | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | V-ATPase / rotary ATPase / complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationproton-transporting V-type ATPase, V1 domain / proton-transporting V-type ATPase, V0 domain / proton-transporting two-sector ATPase complex, catalytic domain / plasma membrane proton-transporting V-type ATPase complex / vacuolar proton-transporting V-type ATPase, V0 domain / vacuolar proton-transporting V-type ATPase, V1 domain / plant-type vacuole / proton-transporting V-type ATPase complex / vacuole / vacuolar acidification ...proton-transporting V-type ATPase, V1 domain / proton-transporting V-type ATPase, V0 domain / proton-transporting two-sector ATPase complex, catalytic domain / plasma membrane proton-transporting V-type ATPase complex / vacuolar proton-transporting V-type ATPase, V0 domain / vacuolar proton-transporting V-type ATPase, V1 domain / plant-type vacuole / proton-transporting V-type ATPase complex / vacuole / vacuolar acidification / vacuolar membrane / ATP metabolic process / H+-transporting two-sector ATPase / phagocytic vesicle / proton-transporting ATPase activity, rotational mechanism / proton-transporting ATP synthase activity, rotational mechanism / transmembrane transport / ATPase binding / lysosomal membrane / ATP hydrolysis activity / ATP binding / membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
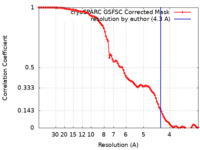
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | |||||||||
 Authors Authors | Abdelaziz RA / Keon KA / Schulze WX / Schumacher K / Rubinstein JL | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Structure of V-ATPase from citrus fruit. Authors: Yong Zi Tan / Kristine A Keon / Rana Abdelaziz / Peter Imming / Waltraud Schulze / Karin Schumacher / John L Rubinstein /   Abstract: We used the Legionella pneumophila effector SidK to affinity purify the endogenous vacuolar-type ATPases (V-ATPases) from lemon fruit. The preparation was sufficient for cryoelectron microscopy, ...We used the Legionella pneumophila effector SidK to affinity purify the endogenous vacuolar-type ATPases (V-ATPases) from lemon fruit. The preparation was sufficient for cryoelectron microscopy, allowing structure determination of the enzyme in two rotational states. The structure defines the ATP:H ratio of the enzyme, demonstrating that it can establish a maximum ΔpH of ∼3, which is insufficient to maintain the low pH observed in the vacuoles of juice sac cells in lemons and other citrus fruit. Compared with yeast and mammalian enzymes, the membrane region of the plant V-ATPase lacks subunit f and possesses an unusual configuration of transmembrane α helices. Subunit H, which inhibits ATP hydrolysis in the isolated catalytic region of V-ATPase, adopts two different conformations in the intact complex, hinting at a role in modulating activity in the intact enzyme. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26826.map.gz emd_26826.map.gz | 94 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26826-v30.xml emd-26826-v30.xml emd-26826.xml emd-26826.xml | 31 KB 31 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26826_fsc.xml emd_26826_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_26826.png emd_26826.png | 61.8 KB | ||
| Filedesc metadata |  emd-26826.cif.gz emd-26826.cif.gz | 8.2 KB | ||
| Others |  emd_26826_additional_1.map.gz emd_26826_additional_1.map.gz emd_26826_half_map_1.map.gz emd_26826_half_map_1.map.gz emd_26826_half_map_2.map.gz emd_26826_half_map_2.map.gz | 97.4 MB 95.6 MB 95.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26826 http://ftp.pdbj.org/pub/emdb/structures/EMD-26826 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26826 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26826 | HTTPS FTP |
-Validation report
| Summary document |  emd_26826_validation.pdf.gz emd_26826_validation.pdf.gz | 902.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26826_full_validation.pdf.gz emd_26826_full_validation.pdf.gz | 901.7 KB | Display | |
| Data in XML |  emd_26826_validation.xml.gz emd_26826_validation.xml.gz | 18.2 KB | Display | |
| Data in CIF |  emd_26826_validation.cif.gz emd_26826_validation.cif.gz | 23.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26826 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26826 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26826 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26826 | HTTPS FTP |
-Related structure data
| Related structure data |  7uwaMC  7uw9C  7uwbC  7uwcC  7uwdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26826.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26826.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Citrus V-ATPase State 1, H in contact with subunits AB | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.617 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Globally refined map
| File | emd_26826_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Globally refined map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Citrus V-ATPase State 1, H in contact with subunits AB
| File | emd_26826_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Citrus V-ATPase State 1, H in contact with subunits AB | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Citrus V-ATPase State 1, H in contact with subunits AB
| File | emd_26826_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Citrus V-ATPase State 1, H in contact with subunits AB | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Citrus V-ATPase
+Supramolecule #1: Citrus V-ATPase
+Macromolecule #1: V-type proton ATPase catalytic subunit A
+Macromolecule #2: V-type proton ATPase subunit B2
+Macromolecule #3: V-type proton ATPase subunit D
+Macromolecule #4: V-type proton ATPase subunit F
+Macromolecule #5: V-type proton ATPase subunit AP1 fragment
+Macromolecule #6: V-type proton ATPase subunit c"
+Macromolecule #7: V-type proton ATPase subunit d2
+Macromolecule #8: V-type proton ATPase subunit c
+Macromolecule #9: V-type proton ATPase subunit AP2 fragment
+Macromolecule #10: V-type proton ATPase subunit E
+Macromolecule #11: V-type proton ATPase subunit G
+Macromolecule #12: V-type proton ATPase subunit C
+Macromolecule #13: V-type proton ATPase subunit a3
+Macromolecule #14: V-type proton ATPase subunit e1
+Macromolecule #15: V-type proton ATPase subunit H
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 31.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.414211 µm / Nominal defocus min: 0.677558 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X