+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2661 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Plasmodium falciparum 80S ribosome | |||||||||
 Map data Map data | Cryo-EM map of the Plasmodium falciparum 80S ribosome | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Plasmodium falciparum 80S ribosome / Cryo-EM | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 3.4 Å | |||||||||
 Authors Authors | Wong W / Bai XC / Brown A / Fernandez IS / Hanssen E / Condron M / Tan YH / Baum J / Scheres SHW | |||||||||
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: Cryo-EM structure of the Plasmodium falciparum 80S ribosome bound to the anti-protozoan drug emetine. Authors: Wilson Wong / Xiao-chen Bai / Alan Brown / Israel S Fernandez / Eric Hanssen / Melanie Condron / Yan Hong Tan / Jake Baum / Sjors H W Scheres /   Abstract: Malaria inflicts an enormous burden on global human health. The emergence of parasite resistance to front-line drugs has prompted a renewed focus on the repositioning of clinically approved drugs as ...Malaria inflicts an enormous burden on global human health. The emergence of parasite resistance to front-line drugs has prompted a renewed focus on the repositioning of clinically approved drugs as potential anti-malarial therapies. Antibiotics that inhibit protein translation are promising candidates for repositioning. We have solved the cryo-EM structure of the cytoplasmic ribosome from the human malaria parasite, Plasmodium falciparum, in complex with emetine at 3.2 Å resolution. Emetine is an anti-protozoan drug used in the treatment of ameobiasis that also displays potent anti-malarial activity. Emetine interacts with the E-site of the ribosomal small subunit and shares a similar binding site with the antibiotic pactamycin, thereby delivering its therapeutic effect by blocking mRNA/tRNA translocation. As the first cryo-EM structure that visualizes an antibiotic bound to any ribosome at atomic resolution, this establishes cryo-EM as a powerful tool for screening and guiding the design of drugs that target parasite translation machinery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2661.map.gz emd_2661.map.gz | 325.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2661-v30.xml emd-2661-v30.xml emd-2661.xml emd-2661.xml | 9.5 KB 9.5 KB | Display Display |  EMDB header EMDB header |
| Images |  cover_image_EMD2661.png cover_image_EMD2661.png | 322.4 KB | ||
| Others |  emd_2661_half_map_1.map.gz emd_2661_half_map_1.map.gz emd_2661_half_map_2.map.gz emd_2661_half_map_2.map.gz | 277.2 MB 277.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2661 http://ftp.pdbj.org/pub/emdb/structures/EMD-2661 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2661 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2661 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2661.map.gz / Format: CCP4 / Size: 339.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2661.map.gz / Format: CCP4 / Size: 339.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of the Plasmodium falciparum 80S ribosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 2661 half map 1.map
| File | emd_2661_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
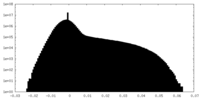
| Density Histograms |
-Supplemental map: emd 2661 half map 2.map
| File | emd_2661_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
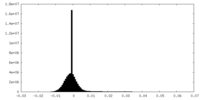
| Density Histograms |
- Sample components
Sample components
-Entire : Plasmodium falciparum 80S ribosome
| Entire | Name: Plasmodium falciparum 80S ribosome |
|---|---|
| Components |
|
-Supramolecule #1000: Plasmodium falciparum 80S ribosome
| Supramolecule | Name: Plasmodium falciparum 80S ribosome / type: sample / ID: 1000 / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 4.2 MDa / Theoretical: 4.2 MDa |
-Supramolecule #1: Plasmodium falciparum 80S ribosome
| Supramolecule | Name: Plasmodium falciparum 80S ribosome / type: complex / ID: 1 / Recombinant expression: No / Ribosome-details: ribosome-eukaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 4.2 MDa / Theoretical: 4.2 MDa |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7.4 Details: 20 mM Hepes pH7.4, 40 mM KCH3COO, 10 mM NH4CH3COO, 10 mM Mg(CH3COO)2 and 5 mM 2-mecaptoethanol |
| Staining | Type: NEGATIVE / Details: Cryo-EM |
| Grid | Details: 30 s on glow-discharged holey carbon grids (Quantifoil R2/2), onto which a home-made continuous carbon film |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 90 K / Instrument: FEI VITROBOT MARK IV / Method: Blot 2.5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 80 K / Max: 90 K / Average: 85 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 78,000 times magnification |
| Date | Jan 28, 2014 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Digitization - Sampling interval: 14 µm / Number real images: 1307 / Average electron dose: 20 e/Å2 Details: An in-house built system was used to intercept the videos from the detector at a rate of 17 frames for the 1 s exposures. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 135922 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.9 µm / Nominal defocus min: 0.7 µm / Nominal magnification: 78000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.4 Å / Resolution method: OTHER / Software - Name: CTFFIND3, RELION Details: Use a newly developed statistical movie processing approach to compensate for beam-induced movement. Number images used: 72293 |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)