+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2632 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Outer arm dynein dimer from mouse respiratory cilia (APO) | |||||||||
 Map data Map data | averaged subtomograms of outer arm dynein dimers from mouse respiratory cilia. Classified as an APO form. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | dynein / cilia / cryo-electron tomography | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 37.0 Å | |||||||||
 Authors Authors | Ueno H / Bui KH / Ishikawa T / Imai Y / Yamaguchi T | |||||||||
 Citation Citation |  Journal: Cytoskeleton (Hoboken) / Year: 2014 Journal: Cytoskeleton (Hoboken) / Year: 2014Title: Structure of dimeric axonemal dynein in cilia suggests an alternative mechanism of force generation. Authors: Hironori Ueno / Khanh Huy Bui / Takuji Ishikawa / Yohsuke Imai / Takami Yamaguchi / Takashi Ishikawa /  Abstract: The mechanism by which the two different heads of the ciliary outer dynein arm produce force to translocate the microtubule during beating is still unknown. In this report we use cryo-electron ...The mechanism by which the two different heads of the ciliary outer dynein arm produce force to translocate the microtubule during beating is still unknown. In this report we use cryo-electron tomography and image processing to analyze the conformational changes and the relative abundance of each conformation of the two dynein heads from mouse respiratory cilia. In the absence of nucleotides the majority of dynein dimers are in the apo form and both heads are tightly packed, whereas they are dissociated and move independently in the presence of nucleotides. The head of the external outer arm dynein heavy chain has a diagonal shift toward both the neighboring B-tubule and the proximal end of the axoneme, while the head of the internal heavy chain shifts only longitudinally toward the proximal end. In the presence of nucleotides a significant number of the dynein dimers have two heads overlapped in the proximal shifting form or overlapped in the apo form. During ciliary bending axonemal dynein translocates microtubules by moving with short steps and two heads stay at the same position longer than cytoplasmic dynein. This demonstrates that the step of the outer arm dynein dimer is not dominated by the hand-over-hand motion, but also indicates the difference between axonemal dynein and cytoplasmic dynein. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2632.map.gz emd_2632.map.gz | 9.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2632-v30.xml emd-2632-v30.xml emd-2632.xml emd-2632.xml | 9.3 KB 9.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2632.tif emd_2632.tif | 759.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2632 http://ftp.pdbj.org/pub/emdb/structures/EMD-2632 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2632 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2632 | HTTPS FTP |
-Validation report
| Summary document |  emd_2632_validation.pdf.gz emd_2632_validation.pdf.gz | 172.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2632_full_validation.pdf.gz emd_2632_full_validation.pdf.gz | 171.6 KB | Display | |
| Data in XML |  emd_2632_validation.xml.gz emd_2632_validation.xml.gz | 4.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2632 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2632 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2632 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2632 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2632.map.gz / Format: CCP4 / Size: 11.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2632.map.gz / Format: CCP4 / Size: 11.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | averaged subtomograms of outer arm dynein dimers from mouse respiratory cilia. Classified as an APO form. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.25 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
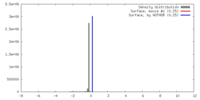
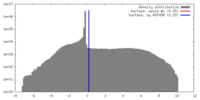
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Outer arm dynein dimer from mouse respiratory cilia in the absenc...
| Entire | Name: Outer arm dynein dimer from mouse respiratory cilia in the absence of additional nucleotides |
|---|---|
| Components |
|
-Supramolecule #1000: Outer arm dynein dimer from mouse respiratory cilia in the absenc...
| Supramolecule | Name: Outer arm dynein dimer from mouse respiratory cilia in the absence of additional nucleotides type: sample / ID: 1000 / Details: in situ sample / Number unique components: 1 |
|---|
-Supramolecule #1: outer arm dyneins with a microtubule doublet
| Supramolecule | Name: outer arm dyneins with a microtubule doublet / type: organelle_or_cellular_component / ID: 1 / Oligomeric state: dimer / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Grid | Details: Quantifoil holey grids |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 100 K / Instrument: FEI VITROBOT MARK IV / Method: Blot for 2 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Min: 90 K / Max: 100 K / Average: 95 K |
| Specialist optics | Energy filter - Name: GIF Tridiem / Energy filter - Lower energy threshold: 20.0 eV / Energy filter - Upper energy threshold: 25.0 eV |
| Date | Nov 9, 2010 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 1000 (2k x 2k) / Average electron dose: 40 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 19303 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.5 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 3.0 µm / Nominal magnification: 30000 |
| Sample stage | Specimen holder: 626 / Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: -60 ° / Tilt series - Axis1 - Max angle: 60 ° |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 37.0 Å / Resolution method: OTHER / Software - Name:  IMOD / Number subtomograms used: 61 IMOD / Number subtomograms used: 61 |
|---|---|
| Final 3D classification | Number classes: 2 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)