+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM maps of the (TGA3)2-(NPR1)2-(TGA3)2 complex | |||||||||
 Map data Map data | consensus refinement map | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
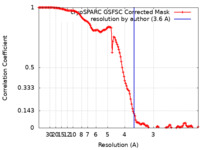
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Wu Q / Zhou Y / Bartesaghi A / Dong X / Zhou P | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Structural basis of NPR1 in activating plant immunity. Authors: Shivesh Kumar / Raul Zavaliev / Qinglin Wu / Ye Zhou / Jie Cheng / Lucas Dillard / Jordan Powers / John Withers / Jinshi Zhao / Ziqiang Guan / Mario J Borgnia / Alberto Bartesaghi / Xinnian Dong / Pei Zhou /  Abstract: NPR1 is a master regulator of the defence transcriptome induced by the plant immune signal salicylic acid. Despite the important role of NPR1 in plant immunity, understanding of its regulatory ...NPR1 is a master regulator of the defence transcriptome induced by the plant immune signal salicylic acid. Despite the important role of NPR1 in plant immunity, understanding of its regulatory mechanisms has been hindered by a lack of structural information. Here we report cryo-electron microscopy and crystal structures of Arabidopsis NPR1 and its complex with the transcription factor TGA3. Cryo-electron microscopy analysis reveals that NPR1 is a bird-shaped homodimer comprising a central Broad-complex, Tramtrack and Bric-à-brac (BTB) domain, a BTB and carboxyterminal Kelch helix bundle, four ankyrin repeats and a disordered salicylic-acid-binding domain. Crystal structure analysis reveals a unique zinc-finger motif in BTB for interacting with ankyrin repeats and mediating NPR1 oligomerization. We found that, after stimulation, salicylic-acid-induced folding and docking of the salicylic-acid-binding domain onto ankyrin repeats is required for the transcriptional cofactor activity of NPR1, providing a structural explanation for a direct role of salicylic acid in regulating NPR1-dependent gene expression. Moreover, our structure of the TGA3-NPR1-TGA3 complex, DNA-binding assay and genetic data show that dimeric NPR1 activates transcription by bridging two fatty-acid-bound TGA3 dimers to form an enhanceosome. The stepwise assembly of the NPR1-TGA complex suggests possible hetero-oligomeric complex formation with other transcription factors, revealing how NPR1 reprograms the defence transcriptome. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25768.map.gz emd_25768.map.gz | 196.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25768-v30.xml emd-25768-v30.xml emd-25768.xml emd-25768.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25768_fsc.xml emd_25768_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_25768.png emd_25768.png | 40.8 KB | ||
| Masks |  emd_25768_msk_1.map emd_25768_msk_1.map | 216 MB |  Mask map Mask map | |
| Others |  emd_25768_additional_1.map.gz emd_25768_additional_1.map.gz emd_25768_additional_2.map.gz emd_25768_additional_2.map.gz emd_25768_additional_3.map.gz emd_25768_additional_3.map.gz | 200.7 MB 200.7 MB 194.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25768 http://ftp.pdbj.org/pub/emdb/structures/EMD-25768 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25768 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25768 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25768.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25768.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | consensus refinement map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.066 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25768_msk_1.map emd_25768_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
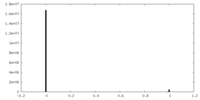
| Density Histograms |
-Additional map: Local Focused Refinement Map (TGA3 dimer)
| File | emd_25768_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local Focused Refinement Map (TGA3 dimer) | ||||||||||||
| Projections & Slices |
| ||||||||||||
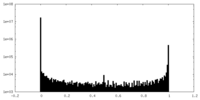
| Density Histograms |
-Additional map: Local Focused Refinement Map (TGA3 dimer)
| File | emd_25768_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local Focused Refinement Map (TGA3 dimer) | ||||||||||||
| Projections & Slices |
| ||||||||||||
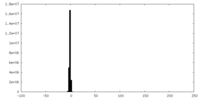
| Density Histograms |
-Additional map: Local Focused Refinement Map (NPR1 dimer)
| File | emd_25768_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local Focused Refinement Map (NPR1 dimer) | ||||||||||||
| Projections & Slices |
| ||||||||||||
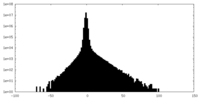
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of (TGA3)2-(NPR1)2-(TGA3)2
| Entire | Name: Complex of (TGA3)2-(NPR1)2-(TGA3)2 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of (TGA3)2-(NPR1)2-(TGA3)2
| Supramolecule | Name: Complex of (TGA3)2-(NPR1)2-(TGA3)2 / type: complex / Chimera: Yes / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.3 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 25 mM HEPES pH7.5, 150 mM NaCl, 2 mM DTT, 0.2 mM salicylic acid |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 277.15 K / Instrument: LEICA EM GP Details: 3 ul of the sample was applied to the grid and incubated for 60 s in the chamber set at 277.15 K and 85% humidity. The grid was blotted for 2.4 s, followed by plunge-freezing in liquid ...Details: 3 ul of the sample was applied to the grid and incubated for 60 s in the chamber set at 277.15 K and 85% humidity. The grid was blotted for 2.4 s, followed by plunge-freezing in liquid ethane cooled by liquid nitrogen.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)