+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25579 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | D3-C3 computationally-designed rotor | |||||||||||||||
 Map data Map data | D3-C3 Computationally-designed Rotor (processed in C3) | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 10.2 Å | |||||||||||||||
 Authors Authors | Hansen JM / Courbet A / Quispe J / Kollman JM / Baker D | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Computational design of mechanically coupled axle-rotor protein assemblies. Authors: A Courbet / J Hansen / Y Hsia / N Bethel / Y-J Park / C Xu / A Moyer / S E Boyken / G Ueda / U Nattermann / D Nagarajan / D-A Silva / W Sheffler / J Quispe / A Nord / N King / P Bradley / D ...Authors: A Courbet / J Hansen / Y Hsia / N Bethel / Y-J Park / C Xu / A Moyer / S E Boyken / G Ueda / U Nattermann / D Nagarajan / D-A Silva / W Sheffler / J Quispe / A Nord / N King / P Bradley / D Veesler / J Kollman / D Baker /    Abstract: Natural molecular machines contain protein components that undergo motion relative to each other. Designing such mechanically constrained nanoscale protein architectures with internal degrees of ...Natural molecular machines contain protein components that undergo motion relative to each other. Designing such mechanically constrained nanoscale protein architectures with internal degrees of freedom is an outstanding challenge for computational protein design. Here we explore the de novo construction of protein machinery from designed axle and rotor components with internal cyclic or dihedral symmetry. We find that the axle-rotor systems assemble in vitro and in vivo as designed. Using cryo-electron microscopy, we find that these systems populate conformationally variable relative orientations reflecting the symmetry of the coupled components and the computationally designed interface energy landscape. These mechanical systems with internal degrees of freedom are a step toward the design of genetically encodable nanomachines. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25579.map.gz emd_25579.map.gz | 102.8 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25579-v30.xml emd-25579-v30.xml emd-25579.xml emd-25579.xml | 14.1 KB 14.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25579.png emd_25579.png | 23.3 KB | ||
| Others |  emd_25579_additional_1.map.gz emd_25579_additional_1.map.gz emd_25579_additional_2.map.gz emd_25579_additional_2.map.gz | 113.1 KB 59.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25579 http://ftp.pdbj.org/pub/emdb/structures/EMD-25579 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25579 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25579 | HTTPS FTP |
-Validation report
| Summary document |  emd_25579_validation.pdf.gz emd_25579_validation.pdf.gz | 290.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25579_full_validation.pdf.gz emd_25579_full_validation.pdf.gz | 290.4 KB | Display | |
| Data in XML |  emd_25579_validation.xml.gz emd_25579_validation.xml.gz | 5.4 KB | Display | |
| Data in CIF |  emd_25579_validation.cif.gz emd_25579_validation.cif.gz | 6.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25579 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25579 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25579 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25579 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25579.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25579.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | D3-C3 Computationally-designed Rotor (processed in C3) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.36 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
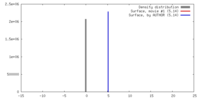
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: D3-C3 Computationally-designed Rotor (processed in D3)
| File | emd_25579_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | D3-C3 Computationally-designed Rotor (processed in D3) | ||||||||||||
| Projections & Slices |
| ||||||||||||
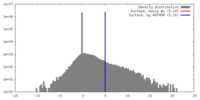
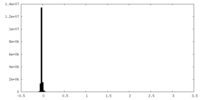
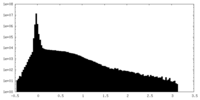
| Density Histograms |
-Additional map: D3-C3 Computationally-designed Rotor (processed in C1)
| File | emd_25579_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | D3-C3 Computationally-designed Rotor (processed in C1) | ||||||||||||
| Projections & Slices |
| ||||||||||||
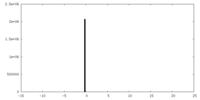
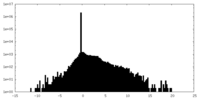
| Density Histograms |
- Sample components
Sample components
-Entire : D3-symmetric axel and C3-symmetric ring
| Entire | Name: D3-symmetric axel and C3-symmetric ring |
|---|---|
| Components |
|
-Supramolecule #1: D3-symmetric axel and C3-symmetric ring
| Supramolecule | Name: D3-symmetric axel and C3-symmetric ring / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 79.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL / In silico model: ab initio |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 10.2 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 16244 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)