+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2509 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of immature HBV core | |||||||||
 Map data Map data | cryo-EM 3-D reconstruction of immature pgRNA-filled HBV core | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HBV / hepatitis B virus / pgRNA | |||||||||
| Biological species |   Hepatitis B virus Hepatitis B virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 14.5 Å | |||||||||
 Authors Authors | Wang JC-Y / Nickens DG / Lentz TB / Loeb DD / Zlotnick A | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2014 Journal: Proc Natl Acad Sci U S A / Year: 2014Title: Encapsidated hepatitis B virus reverse transcriptase is poised on an ordered RNA lattice. Authors: Joseph Che-Yen Wang / David G Nickens / Thomas B Lentz / Daniel D Loeb / Adam Zlotnick /  Abstract: Assembly of a hepatitis B virus (HBV) virion begins with the formation of an RNA-filled core composed of a symmetrical capsid (built of core protein), viral pregenomic RNA, and viral reverse ...Assembly of a hepatitis B virus (HBV) virion begins with the formation of an RNA-filled core composed of a symmetrical capsid (built of core protein), viral pregenomic RNA, and viral reverse transcriptase. To generate the circular dsDNA genome of HBV, reverse transcription requires multiple template switches within the confines of the capsid. To date, most anti-HBV therapeutics target this reverse transcription process. The detailed molecular mechanisms of this crucial process are poorly understood because of the lack of structural information. We hypothesized that capsid, RNA, and viral reverse transcriptase would need a precise geometric organization to accomplish reverse transcription. Here we present the asymmetric structure of authentic RNA-filled cores, determined to 14.5-Å resolution from cryo-EM data. Capsid and RNA are concentric. On the interior of the RNA, we see a distinct donut-like density, assigned to viral reverse transcriptase, which pins the viral pregenomic RNA to the capsid inner surface. The observation of a unique ordered structure inside the core suggests that assembly and the first steps of reverse transcription follow a single, determinate pathway and strongly suggests that all subsequent steps in DNA synthesis do as well. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2509.map.gz emd_2509.map.gz | 6.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2509-v30.xml emd-2509-v30.xml emd-2509.xml emd-2509.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2509.jpg emd_2509.jpg | 100.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2509 http://ftp.pdbj.org/pub/emdb/structures/EMD-2509 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2509 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2509 | HTTPS FTP |
-Validation report
| Summary document |  emd_2509_validation.pdf.gz emd_2509_validation.pdf.gz | 253.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2509_full_validation.pdf.gz emd_2509_full_validation.pdf.gz | 252.8 KB | Display | |
| Data in XML |  emd_2509_validation.xml.gz emd_2509_validation.xml.gz | 5.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2509 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2509 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2509 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2509 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2509.map.gz / Format: CCP4 / Size: 15 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2509.map.gz / Format: CCP4 / Size: 15 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM 3-D reconstruction of immature pgRNA-filled HBV core | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.94 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
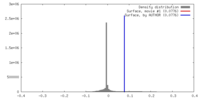
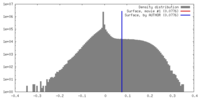
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Immature pgRNA-filled HBV Core
| Entire | Name: Immature pgRNA-filled HBV Core |
|---|---|
| Components |
|
-Supramolecule #1000: Immature pgRNA-filled HBV Core
| Supramolecule | Name: Immature pgRNA-filled HBV Core / type: sample / ID: 1000 / Details: Sample was purified from Huh7-H1 / Number unique components: 1 |
|---|
-Supramolecule #1: Hepatitis B virus
| Supramolecule | Name: Hepatitis B virus / type: virus / ID: 1 / NCBI-ID: 10407 / Sci species name: Hepatitis B virus / Sci species strain: subtype ayw, genotype D / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Host system | Organism:  Homo sapiens (human) / Recombinant cell: Huh7-H1 / Recombinant plasmid: p1159, p1929 Homo sapiens (human) / Recombinant cell: Huh7-H1 / Recombinant plasmid: p1159, p1929 |
| Virus shell | Shell ID: 1 / Name: Cp / Diameter: 340 Å / T number (triangulation number): 4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: 300 mesh continuous carbon film |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 93 K / Instrument: FEI VITROBOT MARK III / Method: Blot for 4 seconds before plunging |
- Electron microscopy #1
Electron microscopy #1
| Microscopy ID | 1 |
|---|---|
| Microscope | JEOL 3200FS |
| Temperature | Average: 97 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 80,000 times magnification |
| Specialist optics | Energy filter - Name: Omega filter / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Dec 20, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 513 / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.1 mm / Nominal magnification: 40000 |
| Sample stage | Specimen holder: 626 / Specimen holder model: GATAN LIQUID NITROGEN |
- Electron microscopy #2
Electron microscopy #2
| Microscopy ID | 2 |
|---|---|
| Microscope | JEOL 3200FS |
| Temperature | Average: 97 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 80,000 times magnification |
| Specialist optics | Energy filter - Name: Omega filter / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Dec 21, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 513 / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.1 mm / Nominal magnification: 40000 |
| Sample stage | Specimen holder: 626 / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Each particle phase-flipping |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 14.5 Å / Resolution method: OTHER / Software - Name: AUTO3DEM / Number images used: 11727 |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)