+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23515 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex | |||||||||||||||||||||
 Map data Map data | XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||
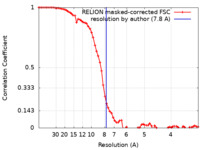
| Method | single particle reconstruction / cryo EM / Resolution: 7.8 Å | |||||||||||||||||||||
 Authors Authors | He Y / Chen S | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Structural basis of long-range to short-range synaptic transition in NHEJ. Authors: Siyu Chen / Linda Lee / Tasmin Naila / Susan Fishbain / Annie Wang / Alan E Tomkinson / Susan P Lees-Miller / Yuan He /   Abstract: DNA double-strand breaks (DSBs) are a highly cytotoxic form of DNA damage and the incorrect repair of DSBs is linked to carcinogenesis. The conserved error-prone non-homologous end joining (NHEJ) ...DNA double-strand breaks (DSBs) are a highly cytotoxic form of DNA damage and the incorrect repair of DSBs is linked to carcinogenesis. The conserved error-prone non-homologous end joining (NHEJ) pathway has a key role in determining the effects of DSB-inducing agents that are used to treat cancer as well as the generation of the diversity in antibodies and T cell receptors. Here we applied single-particle cryo-electron microscopy to visualize two key DNA-protein complexes that are formed by human NHEJ factors. The Ku70/80 heterodimer (Ku), the catalytic subunit of the DNA-dependent protein kinase (DNA-PKcs), DNA ligase IV (LigIV), XRCC4 and XLF form a long-range synaptic complex, in which the DNA ends are held approximately 115 Å apart. Two DNA end-bound subcomplexes comprising Ku and DNA-PKcs are linked by interactions between the DNA-PKcs subunits and a scaffold comprising LigIV, XRCC4, XLF, XRCC4 and LigIV. The relative orientation of the DNA-PKcs molecules suggests a mechanism for autophosphorylation in trans, which leads to the dissociation of DNA-PKcs and the transition into the short-range synaptic complex. Within this complex, the Ku-bound DNA ends are aligned for processing and ligation by the XLF-anchored scaffold, and a single catalytic domain of LigIV is stably associated with a nick between the two Ku molecules, which suggests that the joining of both strands of a DSB involves both LigIV molecules. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23515.map.gz emd_23515.map.gz | 23 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23515-v30.xml emd-23515-v30.xml emd-23515.xml emd-23515.xml | 19.3 KB 19.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23515_fsc.xml emd_23515_fsc.xml | 6.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_23515.png emd_23515.png | 86.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23515 http://ftp.pdbj.org/pub/emdb/structures/EMD-23515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23515 | HTTPS FTP |
-Validation report
| Summary document |  emd_23515_validation.pdf.gz emd_23515_validation.pdf.gz | 315.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23515_full_validation.pdf.gz emd_23515_full_validation.pdf.gz | 314.8 KB | Display | |
| Data in XML |  emd_23515_validation.xml.gz emd_23515_validation.xml.gz | 9.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23515 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23515 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23515 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23515 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_23515.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23515.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.66 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
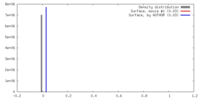
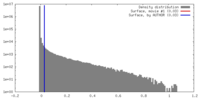
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex
| Entire | Name: XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex |
|---|---|
| Components |
|
-Supramolecule #1: XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex
| Supramolecule | Name: XLF, XRCC4 and LigIV BRCT in NHEJ Short-range synaptic complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: XLF-XRCC4-LigIV BRCT |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 280 KDa |
-Macromolecule #1: XRCC4_HUMAN DNA repair protein XRCC4
| Macromolecule | Name: XRCC4_HUMAN DNA repair protein XRCC4 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
| Sequence | String: MERKISRIHL VSEPSITHFL QVSWEKTLES GFVITLTDGH SAWTGTVSES EISQEADDMA MEKGKYVGEL RKALLSGAGP ADVYTFNFSK ESCYFFFEKN LKDVSFRLGS FNLEKVENPA EVIRELICYC LDTIAENQAK NEHLQKENER LLRDWNDVQG RFEKCVSAKE ...String: MERKISRIHL VSEPSITHFL QVSWEKTLES GFVITLTDGH SAWTGTVSES EISQEADDMA MEKGKYVGEL RKALLSGAGP ADVYTFNFSK ESCYFFFEKN LKDVSFRLGS FNLEKVENPA EVIRELICYC LDTIAENQAK NEHLQKENER LLRDWNDVQG RFEKCVSAKE ALETDLYKRF ILVLNEKKTK IRSLHNKLLN AAQEREKDIK QEGETAICSE MTADRDPVYD ESTDEESENQ TDLSGLASAA VSKDDSIISS LDVTDIAPSR KRRQRMQRNL GTEPKMAPQE NQLQEKENSR PDSSLPETSK KEHISAENMS LETLRNSSPE DLFDEI |
-Macromolecule #2: NHEJ1_HUMAN Non-homologous end-joining factor 1
| Macromolecule | Name: NHEJ1_HUMAN Non-homologous end-joining factor 1 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
| Sequence | String: MEELEQGLLM QPWAWLQLAE NSLLAKVFIT KQGYALLVSD LQQVWHEQVD TSVVSQRAKE LNKRLTAPPA AFLCHLDNLL RPLLKDAAHP SEATFSCDCV ADALILRVRS ELSGLPFYWN FHCMLASPSL VSQHLIRPLM GMSLALQCQV RELATLLHMK DLEIQDYQES ...String: MEELEQGLLM QPWAWLQLAE NSLLAKVFIT KQGYALLVSD LQQVWHEQVD TSVVSQRAKE LNKRLTAPPA AFLCHLDNLL RPLLKDAAHP SEATFSCDCV ADALILRVRS ELSGLPFYWN FHCMLASPSL VSQHLIRPLM GMSLALQCQV RELATLLHMK DLEIQDYQES GATLIRDRLK TEPFEENSFL EQFMIEKLPE ACSIGDGKPF VMNLQDLYMA VTTQEVQVGQ KHQGAGDPHT SNSASLQGID SQCVNQPEQL VSSAPTLSAP EKESTGTSGP LQRPQLSKVK RKKPRGLFS |
-Macromolecule #3: DNLI4_HUMAN DNA ligase 4
| Macromolecule | Name: DNLI4_HUMAN DNA ligase 4 / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
| Sequence | String: SNIFEDVEFC VMSGTDSQPK PDLENRIAEF GGYIVQNPGP DTYCVIAGSE NIRVKNIILS NKHDVVKPAW LLECFKTKSF VPWQPRFMIH MCPSTKEHFA REYDCYGDSY FIDTDLNQLK EVFSGIKNSN EQTPEEMASL IADLEYRYSW DCSPLSMFRR HTVYLDSYAV ...String: SNIFEDVEFC VMSGTDSQPK PDLENRIAEF GGYIVQNPGP DTYCVIAGSE NIRVKNIILS NKHDVVKPAW LLECFKTKSF VPWQPRFMIH MCPSTKEHFA REYDCYGDSY FIDTDLNQLK EVFSGIKNSN EQTPEEMASL IADLEYRYSW DCSPLSMFRR HTVYLDSYAV INDLSTKNEG TRLAIKALEL RFHGAKVVSC LAEGVSHVII GEDHSRVADF KAFRRTFKRK FKILKESWVT DSIDKCELQE ENQYLI |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 Component:
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R3.5/1 / Material: COPPER / Mesh: 200 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Support film - Film thickness: 200.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 101.325 kPa | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 11520 pixel / Digitization - Dimensions - Height: 8184 pixel / Number grids imaged: 2 / Number real images: 32723 / Average exposure time: 0.0426 sec. / Average electron dose: 46.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 30000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Correlation coefficient |
|---|
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)