[English] 日本語
 Yorodumi
Yorodumi- EMDB-22597: Focused cryo-EM map of rabbit RyR1 central and transmembrane doma... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22597 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Focused cryo-EM map of rabbit RyR1 central and transmembrane domains in the presence of AMP-PCP in nanodisc | |||||||||||||||||||||
 Map data Map data | Masked full map | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
| Biological species |   | |||||||||||||||||||||
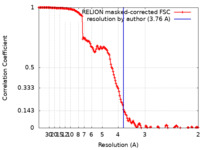
| Method | single particle reconstruction / cryo EM / Resolution: 3.76 Å | |||||||||||||||||||||
 Authors Authors | Nayak AR / Samso M | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Ca inactivation of the mammalian ryanodine receptor type 1 in a lipidic environment revealed by cryo-EM. Authors: Ashok R Nayak / Montserrat Samsó /  Abstract: Activation of the intracellular Ca channel ryanodine receptor (RyR) triggers a cytosolic Ca surge, while elevated cytosolic Ca inhibits the channel in a negative feedback mechanism. Cryogenic ...Activation of the intracellular Ca channel ryanodine receptor (RyR) triggers a cytosolic Ca surge, while elevated cytosolic Ca inhibits the channel in a negative feedback mechanism. Cryogenic electron microscopy of rabbit RyR1 embedded in nanodiscs under partially inactivating Ca conditions revealed an open and a closed-inactivated conformation. Ca binding to the high-affinity site engages the central and C-terminal domains into a block, which pries the S6 four-helix bundle open. Further rotation of this block pushes S6 toward the central axis, closing (inactivating) the channel. Main characteristics of the Ca-inactivated conformation are downward conformation of the cytoplasmic assembly and tightly knit subunit interface contributed by a fully occupied Ca activation site, two inter-subunit resolved lipids, and two salt bridges between the EF hand domain and the S2-S3 loop validated by disease-causing mutations. The structural insight illustrates the prior Ca activation prerequisite for Ca inactivation and provides for a seamless transition from inactivated to closed conformations. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22597.map.gz emd_22597.map.gz | 17.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22597-v30.xml emd-22597-v30.xml emd-22597.xml emd-22597.xml | 25.1 KB 25.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22597_fsc.xml emd_22597_fsc.xml | 15.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_22597.png emd_22597.png | 195.4 KB | ||
| Masks |  emd_22597_msk_1.map emd_22597_msk_1.map | 307.5 MB |  Mask map Mask map | |
| Others |  emd_22597_additional_1.map.gz emd_22597_additional_1.map.gz emd_22597_additional_2.map.gz emd_22597_additional_2.map.gz emd_22597_half_map_1.map.gz emd_22597_half_map_1.map.gz emd_22597_half_map_2.map.gz emd_22597_half_map_2.map.gz | 238.9 MB 239.3 MB 16 MB 16 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22597 http://ftp.pdbj.org/pub/emdb/structures/EMD-22597 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22597 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22597 | HTTPS FTP |
-Related structure data
| Related structure data |  7k0tC  7tdgC  7tdhC  7tdiC  7tdjC  7tdkC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_22597.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22597.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
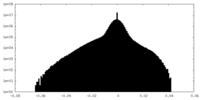
| Annotation | Masked full map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22597_msk_1.map emd_22597_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unmasked and unfiltered half map1 up to Nyquist frequency
| File | emd_22597_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unmasked and unfiltered half map1 up to Nyquist frequency | ||||||||||||
| Projections & Slices |
| ||||||||||||
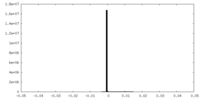
| Density Histograms |
-Additional map: Unmasked and unfiltered half map2 up to Nyquist frequency
| File | emd_22597_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unmasked and unfiltered half map2 up to Nyquist frequency | ||||||||||||
| Projections & Slices |
| ||||||||||||
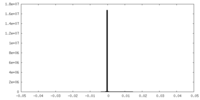
| Density Histograms |
-Half map: Masked half map2
| File | emd_22597_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Masked half map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
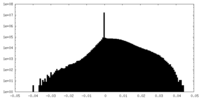
| Density Histograms |
-Half map: Masked half map1
| File | emd_22597_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Masked half map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Rabbit RyR1 with AMP-PCP
| Entire | Name: Rabbit RyR1 with AMP-PCP |
|---|---|
| Components |
|
-Supramolecule #1: Rabbit RyR1 with AMP-PCP
| Supramolecule | Name: Rabbit RyR1 with AMP-PCP / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Purified RyR1 was reconstituted with membrane scaffold protein, MSP1E3D1, and POPC. |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 2.26 MDa |
-Macromolecule #1: RyR1
| Macromolecule | Name: RyR1 / type: protein_or_peptide / ID: 1 Details: Rabbit RyR1 corresponding to amino acid sequence 3668-5037 Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: SFEDRMIDDL SKAGEQEEEE EEVEEKKPDP LHQLVLHFSR TALTEKSKLD EDYLYMAYAD IMAKSCHLEE GGENEEVEV SFEEKEMEKQ RLLYQQSRLH TRGAAEMVLQ MISACKGETG AMVSSTLKLG ISILNGGNAE V QQKMLDYL KDKKEVGFFQ SIQALMQTCS ...String: SFEDRMIDDL SKAGEQEEEE EEVEEKKPDP LHQLVLHFSR TALTEKSKLD EDYLYMAYAD IMAKSCHLEE GGENEEVEV SFEEKEMEKQ RLLYQQSRLH TRGAAEMVLQ MISACKGETG AMVSSTLKLG ISILNGGNAE V QQKMLDYL KDKKEVGFFQ SIQALMQTCS VLDLNAFERQ NKAEGLGMVN EDGTVINRQN GEKVMADDEF TQ DLFRFLQ LLCEGHNNDF QNYLRTQTGN TTTINIIICT VDYLLRLQES ISDFYWYYSG KDVIEEQGKR NFS KAMSVA KQVFNSLTEY IQGPCTGNQQ SLAHSRLWDA VVGFLHVFAH MMMKLAQDSS QIELLKELLD LQKD MVVML LSLLEGNVVN GMIARQMVDM LVESSSNVEM ILKFFDMFLK LKDIVGSEAF QDYVTDPRGL ISKKD FQKA MDSQKQFTGP EIQFLLSCSE ADENEMINFE EFANRFQEPA RDIGFNVAVL LTNLSEHV P HDPRLRNFLE LAESILEYFR PYLGRAEIMG ASRRIERIYF EISETNRAQW EMPQVKESKR QFIFDVVNE GGEAEKMELF VSFCEDTIFE MQIAAQISEF WGELEVQRVK FLNYLSRNFY TLRFLALFLA FAINFILLF YKVSDSPPAN MVYYFLEEST GYMEPALWCL SLLHTLVAFL CIIGYNCLKV PLVIFKREKE LARKLEFDGL YITEQPGDD DVKGQWDRLV LNTPSFPSNY WDKFVKRKVL DKHGDIFGRE RIAELLGMDL ASLEITAHNE R KPDPPPGL LTWLMSIDVK YQIWKFGVIF TDNSFLYLGW YMVMSLLGHY NNFFFAAHLL DIAMGVKTLR TI LSSVTHN GKQLVMTVGL LAVVVYLYTV VAFNFFRKFY NKSEDEDEPD MKCDDMMTCY LFHMYVGVRA GGG IGDEIE DPAGDEYELY RVVFDITFFF FVIVILLAII QGLIIDAFGE LRDQQEQVKE DMETKCFICG IGSD YFDTT PHGFETHTLE EHNLANYMFF LMYLINKDET EHTGQESYVW KMYQERCWDF FPAGDCFRKQ YEDQL S |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.35 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Sample was blotted for 1 second on both sides with Whatman hardened ashless filter paper with blot force 2.. | |||||||||||||||||||||
| Details | Purified RyR1 was reconstituted with membrane scaffold protein, MSP1E3D1, and POPC in 1:2:50 molar ratio. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 1 / Number real images: 1959 / Average electron dose: 70.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)