+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22373 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | RhBZ31.1 Fab in complex with BG505 SOSIP.664 | |||||||||
 Map data Map data | RhBZ31.1 Fab in complex with BG505 SOSIP.664 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 25.0 Å | |||||||||
 Authors Authors | Cottrell CA / Nogal B / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2020 Journal: Cell Rep / Year: 2020Title: Mapping Neutralizing Antibody Epitope Specificities to an HIV Env Trimer in Immunized and in Infected Rhesus Macaques. Authors: Fangzhu Zhao / Collin Joyce / Alison Burns / Bartek Nogal / Christopher A Cottrell / Alejandra Ramos / Trevor Biddle / Matthias Pauthner / Rebecca Nedellec / Huma Qureshi / Rosemarie Mason / ...Authors: Fangzhu Zhao / Collin Joyce / Alison Burns / Bartek Nogal / Christopher A Cottrell / Alejandra Ramos / Trevor Biddle / Matthias Pauthner / Rebecca Nedellec / Huma Qureshi / Rosemarie Mason / Elise Landais / Bryan Briney / Andrew B Ward / Dennis R Burton / Devin Sok /  Abstract: BG505 SOSIP is a well-characterized near-native recombinant HIV Envelope (Env) trimer that holds promise as part of a sequential HIV immunogen regimen to induce broadly neutralizing antibodies (bnAbs) ...BG505 SOSIP is a well-characterized near-native recombinant HIV Envelope (Env) trimer that holds promise as part of a sequential HIV immunogen regimen to induce broadly neutralizing antibodies (bnAbs). Rhesus macaques are considered the most appropriate pre-clinical animal model for monitoring antibody (Ab) responses. Accordingly, we report here the isolation of 45 BG505 autologous neutralizing antibodies (nAbs) with multiple specificities from SOSIP-immunized and BG505 SHIV-infected rhesus macaques. We associate the most potent neutralization with two epitopes: the C3/V5 and V1/V3 regions. We show that all of the nAbs bind in close proximity to known bnAb epitopes and might therefore sterically hinder elicitation of bnAbs. We also identify a "public clonotype" that targets the immunodominant C3/V5 nAb epitope, which suggests that common antibody rearrangements might help determine humoral responses to Env immunogens. The results highlight important considerations for vaccine design in anticipation of results of the BG505 SOSIP trimer in clinical trials. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22373.map.gz emd_22373.map.gz | 20.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22373-v30.xml emd-22373-v30.xml emd-22373.xml emd-22373.xml | 9.8 KB 9.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22373.png emd_22373.png | 44.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22373 http://ftp.pdbj.org/pub/emdb/structures/EMD-22373 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22373 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22373 | HTTPS FTP |
-Validation report
| Summary document |  emd_22373_validation.pdf.gz emd_22373_validation.pdf.gz | 78.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22373_full_validation.pdf.gz emd_22373_full_validation.pdf.gz | 77.2 KB | Display | |
| Data in XML |  emd_22373_validation.xml.gz emd_22373_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22373 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22373 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22373 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22373 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_22373.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22373.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RhBZ31.1 Fab in complex with BG505 SOSIP.664 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
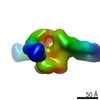
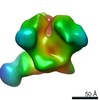
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.98 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
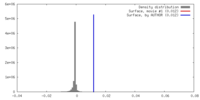
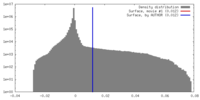
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : RhBZ05.1 Fab in complex with BG505 SOSIP.664
| Entire | Name: RhBZ05.1 Fab in complex with BG505 SOSIP.664 |
|---|---|
| Components |
|
-Supramolecule #1: RhBZ05.1 Fab in complex with BG505 SOSIP.664
| Supramolecule | Name: RhBZ05.1 Fab in complex with BG505 SOSIP.664 / type: complex / ID: 1 / Parent: 0 |
|---|
-Supramolecule #2: BG505 SOSIP.664
| Supramolecule | Name: BG505 SOSIP.664 / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: RhBZ05.1 Fab
| Supramolecule | Name: RhBZ05.1 Fab / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Staining | Type: NEGATIVE / Material: Uranyl formate |
| Grid | Details: unspecified |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI CETA (4k x 4k) / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: RELION (ver. 3.0) / Number images used: 9891 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)