[English] 日本語
 Yorodumi
Yorodumi- EMDB-19283: Conformational Landscape of the Type V-K CRISPR-associated Transp... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Conformational Landscape of the Type V-K CRISPR-associated TransposonIntegration Assembly CAST V-K TnsC domain local-refinement map | |||||||||
 Map data Map data | CAST V-K TnsC domain local-refinement map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR-associated Transposon genome editing transposition / DNA BINDING PROTEIN | |||||||||
| Function / homology | Bacterial TniB / Bacterial TniB protein / : / P-loop containing nucleoside triphosphate hydrolase / TnsC Function and homology information Function and homology information | |||||||||
| Biological species |  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) | |||||||||
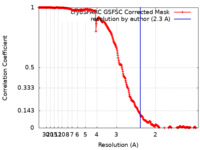
| Method | single particle reconstruction / cryo EM / Resolution: 2.3 Å | |||||||||
 Authors Authors | Tenjo-Castano F / Mesa P / Montoya G | |||||||||
| Funding support |  Denmark, European Union, 2 items Denmark, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2024 Journal: Mol Cell / Year: 2024Title: Conformational landscape of the type V-K CRISPR-associated transposon integration assembly. Authors: Francisco Tenjo-Castaño / Nicholas Sofos / Luisa S Stutzke / Piero Temperini / Anders Fuglsang / Tillmann Pape / Pablo Mesa / Guillermo Montoya /  Abstract: CRISPR-associated transposons (CASTs) are mobile genetic elements that co-opt CRISPR-Cas systems for RNA-guided DNA transposition. CASTs integrate large DNA cargos into the attachment (att) site ...CRISPR-associated transposons (CASTs) are mobile genetic elements that co-opt CRISPR-Cas systems for RNA-guided DNA transposition. CASTs integrate large DNA cargos into the attachment (att) site independently of homology-directed repair and thus hold promise for eukaryotic genome engineering. However, the functional diversity and complexity of CASTs hinder an understanding of their mechanisms. Here, we present the high-resolution cryoelectron microscopy (cryo-EM) structure of the reconstituted ∼1 MDa post-transposition complex of the type V-K CAST, together with different assembly intermediates and diverse TnsC filament lengths, thus enabling the recapitulation of the integration complex formation. The results of mutagenesis experiments probing the roles of specific residues and TnsB-binding sites show that transposition activity can be enhanced and suggest that the distance between the PAM and att sites is determined by the lengths of the TnsB C terminus and the TnsC filament. This singular model of RNA-guided transposition provides a foundation for repurposing the system for genome-editing applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19283.map.gz emd_19283.map.gz | 98.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19283-v30.xml emd-19283-v30.xml emd-19283.xml emd-19283.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19283_fsc.xml emd_19283_fsc.xml | 28.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_19283.png emd_19283.png | 251.9 KB | ||
| Masks |  emd_19283_msk_1.map emd_19283_msk_1.map | 106.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19283.cif.gz emd-19283.cif.gz | 6.5 KB | ||
| Others |  emd_19283_half_map_1.map.gz emd_19283_half_map_1.map.gz emd_19283_half_map_2.map.gz emd_19283_half_map_2.map.gz | 98.6 MB 98.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19283 http://ftp.pdbj.org/pub/emdb/structures/EMD-19283 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19283 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19283 | HTTPS FTP |
-Validation report
| Summary document |  emd_19283_validation.pdf.gz emd_19283_validation.pdf.gz | 1005.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19283_full_validation.pdf.gz emd_19283_full_validation.pdf.gz | 1005.2 KB | Display | |
| Data in XML |  emd_19283_validation.xml.gz emd_19283_validation.xml.gz | 21 KB | Display | |
| Data in CIF |  emd_19283_validation.cif.gz emd_19283_validation.cif.gz | 31.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19283 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19283 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19283 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19283 | HTTPS FTP |
-Related structure data
| Related structure data |  8rkuMC  8axaC  8axbC  8rduC  8rktC  8rkvC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19283.map.gz / Format: CCP4 / Size: 106.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19283.map.gz / Format: CCP4 / Size: 106.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CAST V-K TnsC domain local-refinement map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.728 Å | ||||||||||||||||||||||||||||||||||||
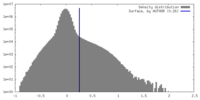
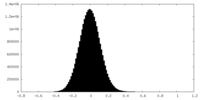
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19283_msk_1.map emd_19283_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
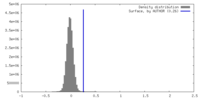
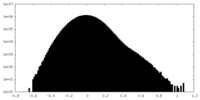
| Density Histograms |
-Half map: CAST V-K TnsC domain local refinement, half-A map
| File | emd_19283_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CAST V-K TnsC domain local refinement, half-A map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: CAST V-K TnsC domain local refinement, half-B map
| File | emd_19283_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CAST V-K TnsC domain local refinement, half-B map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Type V-K CRISPR-associated transposon post-transposition state af...
| Entire | Name: Type V-K CRISPR-associated transposon post-transposition state after transesterification |
|---|---|
| Components |
|
-Supramolecule #1: Type V-K CRISPR-associated transposon post-transposition state af...
| Supramolecule | Name: Type V-K CRISPR-associated transposon post-transposition state after transesterification type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 1 MDa |
-Macromolecule #1: Non-target strand - LE
| Macromolecule | Name: Non-target strand - LE / type: dna / ID: 1 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 20.961504 KDa |
| Sequence | String: (DG)(DT)(DG)(DA)(DA)(DG)(DG)(DT)(DT)(DC) (DT)(DC)(DT)(DT)(DC)(DA)(DG)(DT)(DA)(DT) (DT)(DA)(DA)(DT)(DA)(DA)(DG)(DG)(DC) (DC)(DA)(DC)(DT)(DG)(DT)(DT)(DA)(DA)(DA) (DA) (DC)(DG)(DT)(DA)(DC)(DT) ...String: (DG)(DT)(DG)(DA)(DA)(DG)(DG)(DT)(DT)(DC) (DT)(DC)(DT)(DT)(DC)(DA)(DG)(DT)(DA)(DT) (DT)(DA)(DA)(DT)(DA)(DA)(DG)(DG)(DC) (DC)(DA)(DC)(DT)(DG)(DT)(DT)(DA)(DA)(DA) (DA) (DC)(DG)(DT)(DA)(DC)(DT)(DA)(DT) (DA)(DT)(DA)(DG)(DA)(DC)(DA)(DT)(DC)(DT) (DC)(DC) (DA)(DC)(DA)(DA)(DA)(DA)(DG) (DG) |
-Macromolecule #2: Target strand - LE
| Macromolecule | Name: Target strand - LE / type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 40.82216 KDa |
| Sequence | String: (DA)(DA)(DT)(DT)(DA)(DA)(DA)(DT)(DA)(DG) (DT)(DC)(DA)(DC)(DA)(DA)(DT)(DG)(DA)(DC) (DA)(DT)(DT)(DA)(DA)(DT)(DC)(DT)(DG) (DT)(DC)(DA)(DC)(DC)(DG)(DA)(DC)(DG)(DA) (DC) (DA)(DG)(DA)(DT)(DA)(DA) ...String: (DA)(DA)(DT)(DT)(DA)(DA)(DA)(DT)(DA)(DG) (DT)(DC)(DA)(DC)(DA)(DA)(DT)(DG)(DA)(DC) (DA)(DT)(DT)(DA)(DA)(DT)(DC)(DT)(DG) (DT)(DC)(DA)(DC)(DC)(DG)(DA)(DC)(DG)(DA) (DC) (DA)(DG)(DA)(DT)(DA)(DA)(DT)(DT) (DT)(DG)(DT)(DC)(DA)(DC)(DT)(DG)(DT)(DA) (DC)(DA) (DC)(DT)(DA)(DC)(DG)(DC)(DC) (DT)(DT)(DT)(DT)(DG)(DT)(DG)(DG)(DA)(DG) (DA)(DT)(DG) (DT)(DC)(DT)(DA)(DA)(DT) (DA)(DT)(DC)(DT)(DA)(DC)(DG)(DT)(DT)(DT) (DT)(DA)(DA)(DC) (DA)(DG)(DT)(DG)(DG) (DC)(DC)(DT)(DT)(DA)(DT)(DT)(DA)(DA)(DA) (DT)(DG)(DA)(DC)(DT) (DT)(DC)(DT)(DC) (DA)(DA)(DC)(DC)(DT)(DT)(DC)(DA)(DC) |
-Macromolecule #3: ShTnsC
| Macromolecule | Name: ShTnsC / type: protein_or_peptide / ID: 3 / Number of copies: 14 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 31.400496 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: STEAQAIAKQ LGGVKPDDEW LQAEIARLKG KSIVPLQQVK TLHDWLDGKR KARKSCRVVG ESRTGKTVAC DAYRYRHKPQ QEAGRPPTV PVVYIRPHQK CGPKDLFKKI TEYLKYRVTK GTVSDFRDRT IEVLKGCGVE MLIIDEADRL KPETFADVRD I AEDLGIAV ...String: STEAQAIAKQ LGGVKPDDEW LQAEIARLKG KSIVPLQQVK TLHDWLDGKR KARKSCRVVG ESRTGKTVAC DAYRYRHKPQ QEAGRPPTV PVVYIRPHQK CGPKDLFKKI TEYLKYRVTK GTVSDFRDRT IEVLKGCGVE MLIIDEADRL KPETFADVRD I AEDLGIAV VLVGTDRLDA VIKRDEQVLE RFRAHLRFGK LSGEDFKNTV EMWEQMVLKL PVSSNLKSKE MLRILTSATE GY IGRLDEI LREAAIRSLS RGLKKIDKAV LQEVAKEYK UniProtKB: TnsC |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 14 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 14 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 538 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: UltrAuFoil R0./1 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number real images: 19936 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Protocol: AB INITIO MODEL |
| Output model |  PDB-8rku: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)