+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of NR5A2-nucleosome complex SHL+5.5 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Nucleosome / nuclear receptor / NR5A2 / DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationDeposition of new CENPA-containing nucleosomes at the centromere / Inhibition of DNA recombination at telomere / SUMOylation of chromatin organization proteins / positive regulation of glucocorticoid biosynthetic process / zygotic genome activation / positive regulation of tendon cell differentiation / DNA Damage/Telomere Stress Induced Senescence / morula formation / Regulation of gene expression in early pancreatic precursor cells / G2/M DNA damage checkpoint ...Deposition of new CENPA-containing nucleosomes at the centromere / Inhibition of DNA recombination at telomere / SUMOylation of chromatin organization proteins / positive regulation of glucocorticoid biosynthetic process / zygotic genome activation / positive regulation of tendon cell differentiation / DNA Damage/Telomere Stress Induced Senescence / morula formation / Regulation of gene expression in early pancreatic precursor cells / G2/M DNA damage checkpoint / Negative Regulation of CDH1 Gene Transcription / Regulation of endogenous retroelements by KRAB-ZFP proteins / primary ovarian follicle growth / Condensation of Prophase Chromosomes / HDMs demethylate histones / Nonhomologous End-Joining (NHEJ) / Recognition and association of DNA glycosylase with site containing an affected purine / E3 ubiquitin ligases ubiquitinate target proteins / Cleavage of the damaged purine / HDACs deacetylate histones / pancreas morphogenesis / PRC2 methylates histones and DNA / HATs acetylate histones / Processing of DNA double-strand break ends / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / PKMTs methylate histone lysines / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / acinar cell differentiation / inner cell mass cell differentiation / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / RMTs methylate histone arginines / tissue development / Sertoli cell development / positive regulation of stem cell differentiation / Factors involved in megakaryocyte development and platelet production / Estrogen-dependent gene expression / positive regulation of T cell anergy / embryonic cleavage / exocrine pancreas development / embryo development ending in birth or egg hatching / bile acid metabolic process / cartilage development / negative regulation of chondrocyte differentiation / nucleosomal DNA binding / homeostatic process / calcineurin-mediated signaling / somatic stem cell population maintenance / RNA polymerase II core promoter sequence-specific DNA binding / positive regulation of viral genome replication / protein localization to CENP-A containing chromatin / CENP-A containing nucleosome / hormone-mediated signaling pathway / positive regulation of T cell proliferation / neurogenesis / cholesterol homeostasis / cellular response to leukemia inhibitory factor / transcription coregulator binding / SUMOylation of intracellular receptors / Nuclear Receptor transcription pathway / phospholipid binding / negative regulation of inflammatory response / nuclear receptor activity / RNA polymerase II transcription regulator complex / positive regulation of T cell activation / sequence-specific double-stranded DNA binding / structural constituent of chromatin / nucleosome / nucleosome assembly / chromatin organization / positive regulation of cell growth / DNA-binding transcription activator activity, RNA polymerase II-specific / spermatogenesis / Estrogen-dependent gene expression / sequence-specific DNA binding / DNA-binding transcription factor activity, RNA polymerase II-specific / chromosome, telomeric region / transcription cis-regulatory region binding / RNA polymerase II cis-regulatory region sequence-specific DNA binding / chromatin remodeling / DNA-binding transcription factor activity / protein heterodimerization activity / chromatin binding / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / chromatin / positive regulation of transcription by RNA polymerase II / protein-containing complex / DNA binding / zinc ion binding / nucleoplasm / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
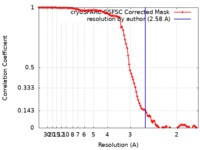
| Method | single particle reconstruction / cryo EM / Resolution: 2.58 Å | |||||||||
 Authors Authors | Kobayashi W / Sappler A / Bollschweiler D / Kummecke M / Basquin J / Arslantas E / Ruangroengkulrith S / Hornberger R / Duderstadt K / Tachibana K | |||||||||
| Funding support |  Germany, European Union, 2 items Germany, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Nucleosome-bound NR5A2 structure reveals pioneer factor mechanism by DNA minor groove anchor competition. Authors: Wataru Kobayashi / Anna H Sappler / Daniel Bollschweiler / Maximilian Kümmecke / Jérôme Basquin / Eda Nur Arslantas / Siwat Ruangroengkulrith / Renate Hornberger / Karl Duderstadt / Kikuë Tachibana /  Abstract: Gene expression during natural and induced reprogramming is controlled by pioneer transcription factors that initiate transcription from closed chromatin. Nr5a2 is a key pioneer factor that regulates ...Gene expression during natural and induced reprogramming is controlled by pioneer transcription factors that initiate transcription from closed chromatin. Nr5a2 is a key pioneer factor that regulates zygotic genome activation in totipotent embryos, pluripotency in embryonic stem cells and metabolism in adult tissues, but the mechanism of its pioneer activity remains poorly understood. Here, we present a cryo-electron microscopy structure of human NR5A2 bound to a nucleosome. The structure shows that the conserved carboxy-terminal extension (CTE) loop of the NR5A2 DNA-binding domain competes with a DNA minor groove anchor of the nucleosome and releases entry-exit site DNA. Mutational analysis showed that NR5A2 D159 of the CTE is dispensable for DNA binding but required for stable nucleosome association and persistent DNA 'unwrapping'. These findings suggest that NR5A2 belongs to an emerging class of pioneer factors that can use DNA minor groove anchor competition to destabilize nucleosomes and facilitate gene expression during reprogramming. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17740.map.gz emd_17740.map.gz | 63 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17740-v30.xml emd-17740-v30.xml emd-17740.xml emd-17740.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17740_fsc.xml emd_17740_fsc.xml | 10.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_17740.png emd_17740.png | 70.2 KB | ||
| Filedesc metadata |  emd-17740.cif.gz emd-17740.cif.gz | 6.5 KB | ||
| Others |  emd_17740_half_map_1.map.gz emd_17740_half_map_1.map.gz emd_17740_half_map_2.map.gz emd_17740_half_map_2.map.gz | 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17740 http://ftp.pdbj.org/pub/emdb/structures/EMD-17740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17740 | HTTPS FTP |
-Related structure data
| Related structure data |  8pkiMC  8pkjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17740.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17740.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8512 Å | ||||||||||||||||||||||||||||||||||||
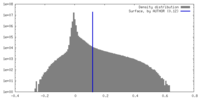
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_17740_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
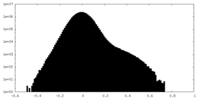
| Density Histograms |
-Half map: #1
| File | emd_17740_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
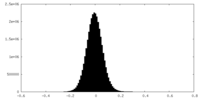
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of the nucleosome containing NR5A2 motif at SHL+5.5
| Entire | Name: Cryo-EM structure of the nucleosome containing NR5A2 motif at SHL+5.5 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of the nucleosome containing NR5A2 motif at SHL+5.5
| Supramolecule | Name: Cryo-EM structure of the nucleosome containing NR5A2 motif at SHL+5.5 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Histone H3.3
| Macromolecule | Name: Histone H3.3 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.360983 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MARTKQTARK STGGKAPRKQ LATKAARKSA PSTGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSAA IGALQEASEA YLVGLFEDTN LCAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.3 |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.394426 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A
| Macromolecule | Name: Histone H2A / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.165551 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKQGGK ARAKAKTRSS RAGLQFPVGR VHRLLRKGNY SERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAIR NDEELNKLLG RVTIAQGGVL PNIQAVLLPK KTESHHKAKG K UniProtKB: Histone H2A |
-Macromolecule #4: Histone H2B type 1-C/E/G
| Macromolecule | Name: Histone H2B type 1-C/E/G / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.937213 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSV YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSSK UniProtKB: Histone H2B type 1-C/E/G |
-Macromolecule #7: Nuclear receptor subfamily 5 group A member 2
| Macromolecule | Name: Nuclear receptor subfamily 5 group A member 2 / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.989939 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LCPVCGDKVS GYHYGLLTCE SCKGFFKRTV QNNKRYTCIE NQNCQIDKTQ RKRCPYCRFQ KCLSVGMKLE AVRADRMRGG RNKFGPMYK RDRAL UniProtKB: Nuclear receptor subfamily 5 group A member 2 |
-Macromolecule #5: DNA
| Macromolecule | Name: DNA / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 46.968922 KDa |
| Sequence | String: (DA)(DT)(DC)(DC)(DT)(DG)(DG)(DA)(DG)(DA) (DA)(DT)(DC)(DC)(DC)(DG)(DG)(DT)(DG)(DC) (DC)(DG)(DA)(DG)(DG)(DC)(DC)(DG)(DC) (DT)(DC)(DA)(DA)(DT)(DT)(DG)(DG)(DT)(DC) (DG) (DT)(DA)(DG)(DA)(DC)(DA) ...String: (DA)(DT)(DC)(DC)(DT)(DG)(DG)(DA)(DG)(DA) (DA)(DT)(DC)(DC)(DC)(DG)(DG)(DT)(DG)(DC) (DC)(DG)(DA)(DG)(DG)(DC)(DC)(DG)(DC) (DT)(DC)(DA)(DA)(DT)(DT)(DG)(DG)(DT)(DC) (DG) (DT)(DA)(DG)(DA)(DC)(DA)(DG)(DC) (DT)(DC)(DT)(DA)(DG)(DC)(DA)(DC)(DC)(DG) (DC)(DT) (DT)(DA)(DA)(DA)(DC)(DG)(DC) (DA)(DC)(DG)(DT)(DA)(DC)(DG)(DC)(DG)(DC) (DT)(DG)(DT) (DC)(DC)(DC)(DC)(DC)(DG) (DC)(DG)(DT)(DT)(DT)(DT)(DA)(DA)(DC)(DC) (DG)(DC)(DC)(DA) (DA)(DG)(DG)(DG)(DG) (DA)(DT)(DT)(DA)(DC)(DT)(DC)(DC)(DC)(DT) (DA)(DG)(DT)(DC)(DT) (DC)(DC)(DA)(DG) (DG)(DC)(DA)(DC)(DG)(DT)(DT)(DC)(DA)(DA) (DG)(DG)(DC)(DC)(DA)(DA) (DT)(DA)(DC) (DA)(DT)(DC)(DC)(DT)(DG)(DT)(DG)(DA)(DT) |
-Macromolecule #6: DNA
| Macromolecule | Name: DNA / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 47.489234 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DC)(DA)(DG)(DG)(DA)(DT) (DG)(DT)(DA)(DT)(DT)(DG)(DG)(DC)(DC)(DT) (DT)(DG)(DA)(DA)(DC)(DG)(DT)(DG)(DC) (DC)(DT)(DG)(DG)(DA)(DG)(DA)(DC)(DT)(DA) (DG) (DG)(DG)(DA)(DG)(DT)(DA) ...String: (DA)(DT)(DC)(DA)(DC)(DA)(DG)(DG)(DA)(DT) (DG)(DT)(DA)(DT)(DT)(DG)(DG)(DC)(DC)(DT) (DT)(DG)(DA)(DA)(DC)(DG)(DT)(DG)(DC) (DC)(DT)(DG)(DG)(DA)(DG)(DA)(DC)(DT)(DA) (DG) (DG)(DG)(DA)(DG)(DT)(DA)(DA)(DT) (DC)(DC)(DC)(DC)(DT)(DT)(DG)(DG)(DC)(DG) (DG)(DT) (DT)(DA)(DA)(DA)(DA)(DC)(DG) (DC)(DG)(DG)(DG)(DG)(DG)(DA)(DC)(DA)(DG) (DC)(DG)(DC) (DG)(DT)(DA)(DC)(DG)(DT) (DG)(DC)(DG)(DT)(DT)(DT)(DA)(DA)(DG)(DC) (DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG)(DA) (DG)(DC)(DT)(DG)(DT)(DC)(DT)(DA)(DC)(DG) (DA)(DC)(DC)(DA)(DA) (DT)(DT)(DG)(DA) (DG)(DC)(DG)(DG)(DC)(DC)(DT)(DC)(DG)(DG) (DC)(DA)(DC)(DC)(DG)(DG) (DG)(DA)(DT) (DT)(DC)(DT)(DC)(DC)(DA)(DG)(DG)(DA)(DT) |
-Macromolecule #8: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 8 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 65.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)