+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Respiratory supercomplex (III2-IV2) from Mycobacterium smegmatis | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | RESPIRATORY SUPERCOMPLEX / MEMBRANE PROTEIN / ACTINOBACTERIA / ELECTRON TRANSPORT | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報aerobic electron transport chain / cytochrome-c oxidase / quinol-cytochrome-c reductase / quinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / superoxide dismutase / superoxide dismutase activity / oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen / ATP synthesis coupled electron transport / respiratory electron transport chain ...aerobic electron transport chain / cytochrome-c oxidase / quinol-cytochrome-c reductase / quinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / superoxide dismutase / superoxide dismutase activity / oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen / ATP synthesis coupled electron transport / respiratory electron transport chain / monooxygenase activity / electron transport chain / 2 iron, 2 sulfur cluster binding / oxidoreductase activity / iron ion binding / copper ion binding / heme binding / membrane / metal ion binding / plasma membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  Mycolicibacterium smegmatis (バクテリア) Mycolicibacterium smegmatis (バクテリア) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.8 Å | |||||||||
 データ登録者 データ登録者 | Kovalova T / Krol S / Sjostrand D / Riepl D / Gamiz-Hernandez A / Brzezinski P / Kaila V / Hogbom M | |||||||||
| 資金援助 |  スウェーデン, 1件 スウェーデン, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2024 ジャーナル: Nat Commun / 年: 2024タイトル: Long-range charge transfer mechanism of the IIIIV mycobacterial supercomplex. 著者: Daniel Riepl / Ana P Gamiz-Hernandez / Terezia Kovalova / Sylwia M Król / Sophie L Mader / Dan Sjöstrand / Martin Högbom / Peter Brzezinski / Ville R I Kaila /  要旨: Aerobic life is powered by membrane-bound redox enzymes that shuttle electrons to oxygen and transfer protons across a biological membrane. Structural studies suggest that these energy-transducing ...Aerobic life is powered by membrane-bound redox enzymes that shuttle electrons to oxygen and transfer protons across a biological membrane. Structural studies suggest that these energy-transducing enzymes operate as higher-order supercomplexes, but their functional role remains poorly understood and highly debated. Here we resolve the functional dynamics of the 0.7 MDa IIIIV obligate supercomplex from Mycobacterium smegmatis, a close relative of M. tuberculosis, the causative agent of tuberculosis. By combining computational, biochemical, and high-resolution (2.3 Å) cryo-electron microscopy experiments, we show how the mycobacterial supercomplex catalyses long-range charge transport from its menaquinol oxidation site to the binuclear active site for oxygen reduction. Our data reveal proton and electron pathways responsible for the charge transfer reactions, mechanistic principles of the quinone catalysis, and how unique molecular adaptations, water molecules, and lipid interactions enable the proton-coupled electron transfer (PCET) reactions. Our combined findings provide a mechanistic blueprint of mycobacterial supercomplexes and a basis for developing drugs against pathogenic bacteria. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_17210.map.gz emd_17210.map.gz | 567.7 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-17210-v30.xml emd-17210-v30.xml emd-17210.xml emd-17210.xml | 42.1 KB 42.1 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_17210.png emd_17210.png | 96 KB | ||
| Filedesc metadata |  emd-17210.cif.gz emd-17210.cif.gz | 10.2 KB | ||
| その他 |  emd_17210_half_map_1.map.gz emd_17210_half_map_1.map.gz emd_17210_half_map_2.map.gz emd_17210_half_map_2.map.gz | 557.6 MB 557.5 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17210 http://ftp.pdbj.org/pub/emdb/structures/EMD-17210 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17210 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17210 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8ovcMC  8ovdC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_17210.map.gz / 形式: CCP4 / 大きさ: 600.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_17210.map.gz / 形式: CCP4 / 大きさ: 600.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.8464 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #2
| ファイル | emd_17210_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
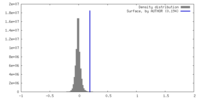
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_17210_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : The respiratory supercomplex
+超分子 #1: The respiratory supercomplex
+分子 #1: Cytochrome bc1 complex cytochrome c subunit
+分子 #2: Cytochrome bc1 complex cytochrome c subunit
+分子 #3: Cytochrome bc1 complex cytochrome b subunit
+分子 #4: Transmembrane protein
+分子 #5: Probable cytochrome c oxidase subunit 3
+分子 #6: Cytochrome c oxidase polypeptide 4
+分子 #7: Cytochrome c oxidase subunit 1
+分子 #8: cytochrome-c oxidase
+分子 #9: Cytochrome c oxidase subunit
+分子 #10: Uncharacterized protein MSMEG_4692/MSMEI_4575
+分子 #11: LpqE protein
+分子 #12: Superoxide dismutase [Cu-Zn]
+分子 #13: Co-purified unknown peptide
+分子 #14: Co-purified unknown peptide
+分子 #15: HEME C
+分子 #16: MENAQUINONE-9
+分子 #17: acyl-phosphatidyl-myo-inositol dimannoside (AcPIM2)
+分子 #18: FE2/S2 (INORGANIC) CLUSTER
+分子 #19: (2R)-2-(hexadecanoyloxy)-3-{[(S)-hydroxy{[(1R,2R,3R,4R,5R,6S)-2,3...
+分子 #20: PROTOPORPHYRIN IX CONTAINING FE
+分子 #21: CARDIOLIPIN
+分子 #22: (1R)-2-(dodecanoyloxy)-1-[(phosphonooxy)methyl]ethyl tetradecanoate
+分子 #23: HEME-A
+分子 #24: COPPER (II) ION
+分子 #25: MAGNESIUM ION
+分子 #26: CALCIUM ION
+分子 #27: (2S)-1-(hexadecanoyloxy)propan-2-yl (10S)-10-methyloctadecanoate
+分子 #28: PALMITIC ACID
+分子 #29: water
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| グリッド | モデル: Quantifoil R2/2 / 材質: COPPER / メッシュ: 300 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 前処理 - タイプ: GLOW DISCHARGE |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK III |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 QUANTUM (4k x 4k) 平均電子線量: 48.2 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 2.2 µm / 最小 デフォーカス(公称値): 0.4 µm / 倍率(公称値): 105000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)