[English] 日本語
 Yorodumi
Yorodumi- EMDB-16925: Focused refinement of HSV-1 DNA polymerase processivity factor in... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Focused refinement of HSV-1 DNA polymerase processivity factor in halted elongation state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA / Polymerase / Complex / TRANSFERASE | |||||||||
| Biological species |   Human alphaherpesvirus 1 strain KOS Human alphaherpesvirus 1 strain KOS | |||||||||
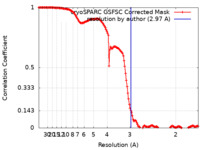
| Method | single particle reconstruction / cryo EM / Resolution: 2.97 Å | |||||||||
 Authors Authors | Gustavsson E / Grunewald K / Elias P / Hallberg M | |||||||||
| Funding support |  Sweden, Sweden,  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2024 Journal: Nucleic Acids Res / Year: 2024Title: Dynamics of the Herpes simplex virus DNA polymerase holoenzyme during DNA synthesis and proof-reading revealed by Cryo-EM. Authors: Emil Gustavsson / Kay Grünewald / Per Elias / B Martin Hällberg /   Abstract: Herpes simplex virus 1 (HSV-1), a double-stranded DNA virus, replicates using seven essential proteins encoded by its genome. Among these, the UL30 DNA polymerase, complexed with the UL42 ...Herpes simplex virus 1 (HSV-1), a double-stranded DNA virus, replicates using seven essential proteins encoded by its genome. Among these, the UL30 DNA polymerase, complexed with the UL42 processivity factor, orchestrates leading and lagging strand replication of the 152 kb viral genome. UL30 polymerase is a prime target for antiviral therapy, and resistance to current drugs can arise in immunocompromised individuals. Using electron cryo-microscopy (cryo-EM), we unveil the dynamic changes of the UL30/UL42 complex with DNA in three distinct states. First, a pre-translocation state with an open fingers domain ready for nucleotide incorporation. Second, a halted elongation state where the fingers close, trapping dATP in the dNTP pocket. Third, a DNA-editing state involving significant conformational changes to allow DNA realignment for exonuclease activity. Additionally, the flexible UL30 C-terminal domain interacts with UL42, forming an extended positively charged surface binding to DNA, thereby enhancing processive synthesis. These findings highlight substantial structural shifts in the polymerase and its DNA interactions during replication, offering insights for future antiviral drug development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16925.map.gz emd_16925.map.gz | 154.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16925-v30.xml emd-16925-v30.xml emd-16925.xml emd-16925.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16925_fsc.xml emd_16925_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_16925.png emd_16925.png | 111.2 KB | ||
| Masks |  emd_16925_msk_1.map emd_16925_msk_1.map | 307.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16925.cif.gz emd-16925.cif.gz | 4.5 KB | ||
| Others |  emd_16925_half_map_1.map.gz emd_16925_half_map_1.map.gz emd_16925_half_map_2.map.gz emd_16925_half_map_2.map.gz | 285 MB 285 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16925 http://ftp.pdbj.org/pub/emdb/structures/EMD-16925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16925 | HTTPS FTP |
-Validation report
| Summary document |  emd_16925_validation.pdf.gz emd_16925_validation.pdf.gz | 1014.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16925_full_validation.pdf.gz emd_16925_full_validation.pdf.gz | 1014.2 KB | Display | |
| Data in XML |  emd_16925_validation.xml.gz emd_16925_validation.xml.gz | 23.2 KB | Display | |
| Data in CIF |  emd_16925_validation.cif.gz emd_16925_validation.cif.gz | 30.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16925 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16925 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16925 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16925 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16925.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16925.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16925_msk_1.map emd_16925_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16925_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16925_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HSV-1 DNA polymerase-processivity factor complex in halted elonga...
| Entire | Name: HSV-1 DNA polymerase-processivity factor complex in halted elongation state |
|---|---|
| Components |
|
-Supramolecule #1: HSV-1 DNA polymerase-processivity factor complex in halted elonga...
| Supramolecule | Name: HSV-1 DNA polymerase-processivity factor complex in halted elongation state type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:   Human alphaherpesvirus 1 strain KOS Human alphaherpesvirus 1 strain KOS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
Details: 20mM HEPES pH7.8, 150mM NaCl, 5mM MgCl2, 2mM DTT | |||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY | |||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)