+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the Fmoc-Tau-PAM4 Type 4 amyloid fibril | |||||||||
 Map data Map data | Final postprocessed, sharpened, helical symmetrised map of the Fmoc-TauPAM4 fibril morphology IIb | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | amyloid / tau / helical / cross-beta / fibril / neurodegeneration / Fmoc / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationplus-end-directed organelle transport along microtubule / axonal transport / histone-dependent DNA binding / neurofibrillary tangle assembly / positive regulation of diacylglycerol kinase activity / negative regulation of establishment of protein localization to mitochondrion / neurofibrillary tangle / positive regulation of protein localization to synapse / microtubule lateral binding / tubulin complex ...plus-end-directed organelle transport along microtubule / axonal transport / histone-dependent DNA binding / neurofibrillary tangle assembly / positive regulation of diacylglycerol kinase activity / negative regulation of establishment of protein localization to mitochondrion / neurofibrillary tangle / positive regulation of protein localization to synapse / microtubule lateral binding / tubulin complex / phosphatidylinositol bisphosphate binding / main axon / regulation of long-term synaptic depression / negative regulation of kinase activity / negative regulation of tubulin deacetylation / generation of neurons / positive regulation of protein localization / rRNA metabolic process / internal protein amino acid acetylation / regulation of chromosome organization / regulation of mitochondrial fission / intracellular distribution of mitochondria / axonal transport of mitochondrion / axon development / central nervous system neuron development / regulation of microtubule polymerization / microtubule polymerization / minor groove of adenine-thymine-rich DNA binding / lipoprotein particle binding / dynactin binding / apolipoprotein binding / glial cell projection / negative regulation of mitochondrial membrane potential / protein polymerization / axolemma / negative regulation of mitochondrial fission / Caspase-mediated cleavage of cytoskeletal proteins / regulation of microtubule polymerization or depolymerization / positive regulation of axon extension / regulation of microtubule cytoskeleton organization / Activation of AMPK downstream of NMDARs / supramolecular fiber organization / regulation of cellular response to heat / stress granule assembly / cytoplasmic microtubule organization / regulation of calcium-mediated signaling / axon cytoplasm / positive regulation of microtubule polymerization / somatodendritic compartment / synapse assembly / cellular response to brain-derived neurotrophic factor stimulus / phosphatidylinositol binding / nuclear periphery / cellular response to nerve growth factor stimulus / positive regulation of superoxide anion generation / protein phosphatase 2A binding / regulation of autophagy / astrocyte activation / response to lead ion / microglial cell activation / synapse organization / Hsp90 protein binding / regulation of synaptic plasticity / PKR-mediated signaling / protein homooligomerization / memory / microtubule cytoskeleton organization / cellular response to reactive oxygen species / SH3 domain binding / cytoplasmic ribonucleoprotein granule / activation of cysteine-type endopeptidase activity involved in apoptotic process / microtubule cytoskeleton / neuron projection development / cell-cell signaling / protein-macromolecule adaptor activity / actin binding / single-stranded DNA binding / cellular response to heat / protein-folding chaperone binding / cell body / growth cone / microtubule binding / double-stranded DNA binding / microtubule / amyloid fibril formation / sequence-specific DNA binding / dendritic spine / learning or memory / nuclear speck / neuron projection / membrane raft / axon / negative regulation of gene expression / neuronal cell body / dendrite / DNA damage response / protein kinase binding / enzyme binding / mitochondrion / DNA binding Similarity search - Function | |||||||||
| Biological species | synthetic construct (others) | |||||||||
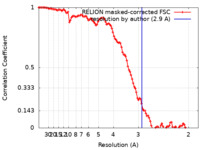
| Method | helical reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Wilkinson M / Louros N / Tsaka G / Ramakers M / Morelli C / Garcia T / Gallardo RU / D'Haeyer S / Goossens V / Audenaert D ...Wilkinson M / Louros N / Tsaka G / Ramakers M / Morelli C / Garcia T / Gallardo RU / D'Haeyer S / Goossens V / Audenaert D / Thal DR / Ranson NA / Radford SE / Rousseau F / Schymkowitz J | |||||||||
| Funding support |  Belgium, 1 items Belgium, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Local structural preferences in shaping tau amyloid polymorphism. Authors: Nikolaos Louros / Martin Wilkinson / Grigoria Tsaka / Meine Ramakers / Chiara Morelli / Teresa Garcia / Rodrigo Gallardo / Sam D'Haeyer / Vera Goossens / Dominique Audenaert / Dietmar Rudolf ...Authors: Nikolaos Louros / Martin Wilkinson / Grigoria Tsaka / Meine Ramakers / Chiara Morelli / Teresa Garcia / Rodrigo Gallardo / Sam D'Haeyer / Vera Goossens / Dominique Audenaert / Dietmar Rudolf Thal / Ian R Mackenzie / Rosa Rademakers / Neil A Ranson / Sheena E Radford / Frederic Rousseau / Joost Schymkowitz /    Abstract: Tauopathies encompass a group of neurodegenerative disorders characterised by diverse tau amyloid fibril structures. The persistence of polymorphism across tauopathies suggests that distinct ...Tauopathies encompass a group of neurodegenerative disorders characterised by diverse tau amyloid fibril structures. The persistence of polymorphism across tauopathies suggests that distinct pathological conditions dictate the adopted polymorph for each disease. However, the extent to which intrinsic structural tendencies of tau amyloid cores contribute to fibril polymorphism remains uncertain. Using a combination of experimental approaches, we here identify a new amyloidogenic motif, PAM4 (Polymorphic Amyloid Motif of Repeat 4), as a significant contributor to tau polymorphism. Calculation of per-residue contributions to the stability of the fibril cores of different pathologic tau structures suggests that PAM4 plays a central role in preserving structural integrity across amyloid polymorphs. Consistent with this, cryo-EM structural analysis of fibrils formed from a synthetic PAM4 peptide shows that the sequence adopts alternative structures that closely correspond to distinct disease-associated tau strains. Furthermore, in-cell experiments revealed that PAM4 deletion hampers the cellular seeding efficiency of tau aggregates extracted from Alzheimer's disease, corticobasal degeneration, and progressive supranuclear palsy patients, underscoring PAM4's pivotal role in these tauopathies. Together, our results highlight the importance of the intrinsic structural propensity of amyloid core segments to determine the structure of tau in cells, and in propagating amyloid structures in disease. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16886.map.gz emd_16886.map.gz | 20.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16886-v30.xml emd-16886-v30.xml emd-16886.xml emd-16886.xml | 19.3 KB 19.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16886_fsc.xml emd_16886_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16886.png emd_16886.png | 78.7 KB | ||
| Filedesc metadata |  emd-16886.cif.gz emd-16886.cif.gz | 6.1 KB | ||
| Others |  emd_16886_half_map_1.map.gz emd_16886_half_map_1.map.gz emd_16886_half_map_2.map.gz emd_16886_half_map_2.map.gz | 81.1 MB 81.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16886 http://ftp.pdbj.org/pub/emdb/structures/EMD-16886 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16886 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16886 | HTTPS FTP |
-Validation report
| Summary document |  emd_16886_validation.pdf.gz emd_16886_validation.pdf.gz | 1017.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16886_full_validation.pdf.gz emd_16886_full_validation.pdf.gz | 1016.8 KB | Display | |
| Data in XML |  emd_16886_validation.xml.gz emd_16886_validation.xml.gz | 18 KB | Display | |
| Data in CIF |  emd_16886_validation.cif.gz emd_16886_validation.cif.gz | 23.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16886 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16886 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16886 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16886 | HTTPS FTP |
-Related structure data
| Related structure data |  8oi0MC  8oh2C  8ohiC  8ohpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16886.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16886.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final postprocessed, sharpened, helical symmetrised map of the Fmoc-TauPAM4 fibril morphology IIb | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.94 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: halfmap1
| File | emd_16886_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap2
| File | emd_16886_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Amyloid fibril form 2b of Tau-PAM4 peptide adducted with the Fmoc...
| Entire | Name: Amyloid fibril form 2b of Tau-PAM4 peptide adducted with the Fmoc protection group |
|---|---|
| Components |
|
-Supramolecule #1: Amyloid fibril form 2b of Tau-PAM4 peptide adducted with the Fmoc...
| Supramolecule | Name: Amyloid fibril form 2b of Tau-PAM4 peptide adducted with the Fmoc protection group type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Synthesised peptide assembled into amyloid fibril |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: Microtubule-associated protein tau
| Macromolecule | Name: Microtubule-associated protein tau / type: protein_or_peptide / ID: 1 Details: 13-residue peptide of the PAM4 motif of Tau, corresponding to residues 350-362 of the Tau repeat domain. The peptide is C-terminally amidated and should be N-terminally acetylated, however ...Details: 13-residue peptide of the PAM4 motif of Tau, corresponding to residues 350-362 of the Tau repeat domain. The peptide is C-terminally amidated and should be N-terminally acetylated, however 10% (by mass spec) still adducted to Fmoc protection group used in synthesis. the Fmoc-peptide form dominated the fibril assembly. Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 1.652269 KDa |
| Sequence | String: (VP1)VQSKIGSLD NIT(HIA) UniProtKB: Microtubule-associated protein tau |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7 Details: Peptide resuspended in MilliQ water for aggregation reaction |
| Grid | Model: EMS Lacey Carbon / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 1957 / Average exposure time: 5.0 sec. / Average electron dose: 32.0 e/Å2 Details: Movies were collected as 1204 EER frames compressed and re-grouped into 35 TIF fractions |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Details | Initial model edited in coot | ||||||
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 64 / Target criteria: Cross-correlation coefficient | ||||||
| Output model |  PDB-8oi0: |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X