[English] 日本語
 Yorodumi
Yorodumi- EMDB-1659: Solution structure and characterisation of the human pyruvate deh... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1659 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Solution structure and characterisation of the human pyruvate dehydrogenase complex core assembly | |||||||||
 Map data Map data | This contour level displays the E2-E3BP-E3 structure with ordered extensions from the core surface and some noisier density not following the icosahedral symmetry | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Pyruvate dehydrogenase complex / dodecahedron | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 33.0 Å | |||||||||
 Authors Authors | Vijayakrishnan S / Kelly SM / Gilbert RJC / Callow P / Bhella D / Forsyth T / Lindsay JG / Byron O | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2010 Journal: J Mol Biol / Year: 2010Title: Solution structure and characterisation of the human pyruvate dehydrogenase complex core assembly. Authors: S Vijayakrishnan / S M Kelly / R J C Gilbert / P Callow / D Bhella / T Forsyth / J G Lindsay / O Byron /  Abstract: Mammalian pyruvate dehydrogenase complex (PDC) is a key multi-enzyme assembly that is responsible for glucose homeostasis maintenance and conversion of pyruvate into acetyl-CoA. It comprises a ...Mammalian pyruvate dehydrogenase complex (PDC) is a key multi-enzyme assembly that is responsible for glucose homeostasis maintenance and conversion of pyruvate into acetyl-CoA. It comprises a central pentagonal dodecahedral core consisting of two subunit types (E2 and E3BP) to which peripheral enzymes (E1 and E3) bind tightly but non-covalently. Currently, there are two conflicting models of PDC (E2+E3BP) core organisation: the 'addition' model (60+12) and the 'substitution' model (48+12). Here we present the first ever low-resolution structures of human recombinant full-length PDC core (rE2/E3BP), truncated PDC core (tE2/E3BP) and native bovine heart PDC core (bE2/E3BP) obtained by small-angle X-ray scattering and small-angle neutron scattering. These structures, corroborated by negative-stain and cryo electron microscopy data, clearly reveal open pentagonal core faces, favouring the 'substitution' model of core organisation. The native and recombinant core structures are all similar to the truncated bacterial E2 core crystal structure obtained previously. Cryo-electron microscopy reconstructions of rE2/E3BP and rE2/E3BP:E3 directly confirm that the core has open pentagonal faces, agree with scattering-derived models and show density extending outwards from their surfaces, which is much more structurally ordered in the presence of E3. Additionally, analytical ultracentrifugation characterisation of rE2/E3BP, rE2 (full-length recombinant E2-only) and tE2/E3BP supports the substitution model. Superimposition of the small-angle neutron scattering tE2/E3BP and truncated bacterial E2 crystal structures demonstrates conservation of the overall pentagonal dodecahedral morphology, despite evolutionary diversity. In addition, unfolding studies using circular dichroism and tryptophan fluorescence spectroscopy show that the rE2/E3BP is less stable than its rE2 counterpart, indicative of a role for E3BP in core destabilisation. The architectural complexity and lower stability of the E2/E3BP core may be of benefit to mammals, where sophisticated fine-tuning is required for cores with optimal catalytic and regulatory efficiencies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1659.map.gz emd_1659.map.gz | 28.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1659-v30.xml emd-1659-v30.xml emd-1659.xml emd-1659.xml | 12.6 KB 12.6 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-1659.gif EMD-1659.gif | 129.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1659 http://ftp.pdbj.org/pub/emdb/structures/EMD-1659 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1659 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1659 | HTTPS FTP |
-Validation report
| Summary document |  emd_1659_validation.pdf.gz emd_1659_validation.pdf.gz | 240.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1659_full_validation.pdf.gz emd_1659_full_validation.pdf.gz | 239.6 KB | Display | |
| Data in XML |  emd_1659_validation.xml.gz emd_1659_validation.xml.gz | 6.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1659 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1659 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1659 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1659 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1659.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1659.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This contour level displays the E2-E3BP-E3 structure with ordered extensions from the core surface and some noisier density not following the icosahedral symmetry | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
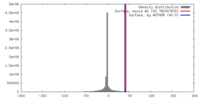
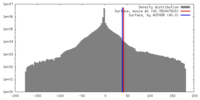
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Pyruvate dehydrogenase E2-E3BP-E3 complex
| Entire | Name: Pyruvate dehydrogenase E2-E3BP-E3 complex |
|---|---|
| Components |
|
-Supramolecule #1000: Pyruvate dehydrogenase E2-E3BP-E3 complex
| Supramolecule | Name: Pyruvate dehydrogenase E2-E3BP-E3 complex / type: sample / ID: 1000 Oligomeric state: A dodecahedral lattice formed of mixed trimers of E2 and E3BP, most likely each containing 2 E2 and 1 E3BP, bound with the E3 Number unique components: 3 |
|---|---|
| Molecular weight | Theoretical: 5.4 MDa |
-Macromolecule #1: dihydrolipoamide dehydrogenase
| Macromolecule | Name: dihydrolipoamide dehydrogenase / type: protein_or_peptide / ID: 1 / Name.synonym: E3 / Number of copies: 20 / Oligomeric state: Hetero-icosahedron / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 60 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: E3 binding protein
| Macromolecule | Name: E3 binding protein / type: protein_or_peptide / ID: 2 / Name.synonym: E3BP / Number of copies: 20 / Oligomeric state: Hetero-icosahedron / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 60 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #3: Dihydrolipoyl transacetylase
| Macromolecule | Name: Dihydrolipoyl transacetylase / type: protein_or_peptide / ID: 3 / Name.synonym: E2 / Number of copies: 40 / Oligomeric state: Hetero-icosahedron / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 70 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 2 mM EDTA, 0.01% (w/v) NaN3, 100 mM NaCl, 50 mM KH2PO4, pH 7.5 |
|---|---|
| Grid | Details: 300 mesh copper grid with lacey carbon film |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 100 K / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: Homemade plunger Method: Blot briefly before plunging, using Whatman no.1 paper |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Temperature | Average: 115 K |
| Alignment procedure | Legacy - Astigmatism: At 200,000 magnification |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 9 Details: Images were binned from a scanned pixel size at the specimen of 1.8A to 3.6A using Proc2D Od range: 5 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 38000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Particles selected interactively using Boxer |
|---|---|
| CTF correction | Details: Per micrograph |
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 33.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider, WellMAP Details: Final map had converged to given resolution, on a 1 degree spacing. Number images used: 172 |
| Final angle assignment | Details: SPIDER convention |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - #0 - Chain ID: A / Chain - #1 - Chain ID: B / Chain - #2 - Chain ID: C / Chain - #3 - Chain ID: D / Chain - #4 - Chain ID: E |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | PDBEntryID_givenInChain. Protocol: Rigid body, after icosahedral symmetrization. The icosahedral assembly was constructed and then fitted as a rigid body. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: CCC |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)