[English] 日本語
 Yorodumi
Yorodumi- EMDB-15519: human MutSalpha (MSH2/MSH6) on DNA containing a GT mismatch in th... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human MutSalpha (MSH2/MSH6) on DNA containing a GT mismatch in the presence of ADP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Bruekner SR / Liaci AM / Sixma TK | |||||||||
| Funding support |  Netherlands, European Union, 2 items Netherlands, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: Unexpected moves: a conformational change in MutSα enables high-affinity DNA mismatch binding. Authors: Susanne R Bruekner / Wietske Pieters / Alexander Fish / A Manuel Liaci / Serge Scheffers / Emily Rayner / Daphne Kaldenbach / Lisa Drost / Marleen Dekker / Sandrine van Hees-Stuivenberg / ...Authors: Susanne R Bruekner / Wietske Pieters / Alexander Fish / A Manuel Liaci / Serge Scheffers / Emily Rayner / Daphne Kaldenbach / Lisa Drost / Marleen Dekker / Sandrine van Hees-Stuivenberg / Elly Delzenne-Goette / Charlotte de Konink / Hellen Houlleberghs / Hendrikus Jan Dubbink / Abeer AlSaegh / Niels de Wind / Friedrich Förster / Hein Te Riele / Titia K Sixma /  Abstract: The DNA mismatch repair protein MutSα recognizes wrongly incorporated DNA bases and initiates their correction during DNA replication. Dysfunctions in mismatch repair lead to a predisposition to ...The DNA mismatch repair protein MutSα recognizes wrongly incorporated DNA bases and initiates their correction during DNA replication. Dysfunctions in mismatch repair lead to a predisposition to cancer. Here, we study the homozygous mutation V63E in MSH2 that was found in the germline of a patient with suspected constitutional mismatch repair deficiency syndrome who developed colorectal cancer before the age of 30. Characterization of the mutant in mouse models, as well as slippage and repair assays, shows a mildly pathogenic phenotype. Using cryogenic electron microscopy and surface plasmon resonance, we explored the mechanistic effect of this mutation on MutSα function. We discovered that V63E disrupts a previously unappreciated interface between the mismatch binding domains (MBDs) of MSH2 and MSH6 and leads to reduced DNA binding. Our research identifies this interface as a 'safety lock' that ensures high-affinity DNA binding to increase replication fidelity. Our mechanistic model explains the hypomorphic phenotype of the V63E patient mutation and other variants in the MBD interface. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15519.map.gz emd_15519.map.gz | 6.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15519-v30.xml emd-15519-v30.xml emd-15519.xml emd-15519.xml | 25.4 KB 25.4 KB | Display Display |  EMDB header EMDB header |
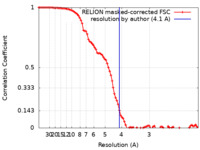
| FSC (resolution estimation) |  emd_15519_fsc.xml emd_15519_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_15519.png emd_15519.png | 157.5 KB | ||
| Others |  emd_15519_half_map_1.map.gz emd_15519_half_map_1.map.gz emd_15519_half_map_2.map.gz emd_15519_half_map_2.map.gz | 49.7 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15519 http://ftp.pdbj.org/pub/emdb/structures/EMD-15519 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15519 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15519 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15519.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15519.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.041 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_15519_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_15519_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : MutSalpha on mismatched DNA in the presence of ADP
| Entire | Name: MutSalpha on mismatched DNA in the presence of ADP |
|---|---|
| Components |
|
-Supramolecule #1: MutSalpha on mismatched DNA in the presence of ADP
| Supramolecule | Name: MutSalpha on mismatched DNA in the presence of ADP / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 31 KDa |
-Supramolecule #2: MSH2
| Supramolecule | Name: MSH2 / type: complex / ID: 2 / Chimera: Yes / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: MSH6
| Supramolecule | Name: MSH6 / type: complex / ID: 3 / Chimera: Yes / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: DNA (50-mer)
| Supramolecule | Name: DNA (50-mer) / type: complex / ID: 4 / Chimera: Yes / Parent: 1 / Macromolecule list: #3-#4 / Details: contains a GT mismatch at position 25 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: MSH2
| Macromolecule | Name: MSH2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MAVQPKETLQ LESAAEVGFV RFFQGMPEKP TTTVRLFDRG DFYTAHGEDA LLAAREVFKT QGVIKYMGPA GAKNLQSVVL SKMNFESFVK DLLLVRQYRV EVYKNRAGNK ASKENDWYLA YKASPGNLSQ FEDILFGNND MSASIGVVGV KMSAVDGQRQ VGVGYVDSIQ ...String: MAVQPKETLQ LESAAEVGFV RFFQGMPEKP TTTVRLFDRG DFYTAHGEDA LLAAREVFKT QGVIKYMGPA GAKNLQSVVL SKMNFESFVK DLLLVRQYRV EVYKNRAGNK ASKENDWYLA YKASPGNLSQ FEDILFGNND MSASIGVVGV KMSAVDGQRQ VGVGYVDSIQ RKLGLCEFPD NDQFSNLEAL LIQIGPKECV LPGGETAGDM GKLRQIIQRG GILITERKKA DFSTKDIYQD LNRLLKGKKG EQMNSAVLPE MENQVAVSSL SAVIKFLELL SDDSNFGQFE LTTFDFSQYM KLDIAAVRAL NLFQGSVEDT TGSQSLAALL NKCKTPQGQR LVNQWIKQPL MDKNRIEERL NLVEAFVEDA ELRQTLQEDL LRRFPDLNRL AKKFQRQAAN LQDCYRLYQG INQLPNVIQA LEKHEGKHQK LLLAVFVTPL TDLRSDFSKF QEMIETTLDM DQVENHEFLV KPSFDPNLSE LREIMNDLEK KMQSTLISAA RDLGLDPGKQ IKLDSSAQFG YYFRVTCKEE KVLRNNKNFS TVDIQKNGVK FTNSKLTSLN EEYTKNKTEY EEAQDAIVKE IVNISSGYVE PMQTLNDVLA QLDAVVSFAH VSNGAPVPYV RPAILEKGQG RIILKASRHA CVEVQDEIAF IPNDVYFEKD KQMFHIITGP NMGGKSTYIR QTGVIVLMAQ IGCFVPCESA EVSIVDCILA RVGAGDSQLK GVSTFMAEML ETASILRSAT KDSLIIIDEL GRGTSTYDGF GLAWAISEYI ATKIGAFCMF ATHFHELTAL ANQIPTVNNL HVTALTTEET LTMLYQVKKG VCDQSFGIHV AELANFPKHV IECAKQKALE LEEFQYIGES QGYDIMEPAA KKCYLEREQG EKIIQEFLSK VKQMPFTEMS EENITIKLKQ LKAEVIAKNN SFVNEIISRI KVTT |
-Macromolecule #2: MSH6
| Macromolecule | Name: MSH6 / type: protein_or_peptide / ID: 2 / Details: contains an N-terminal 10His-TwinStrep tag / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MAHHHHHHHH HHGSAASWSH PQFEKGGGSG GGSGGGSWSH PQFEKGALEV LFQGPSRQST LYSFFPKSPA LSDANKASAR ASREGGRAAA APEASPSPGG DAAWSEAGPG PRPLARSASP PKAKNLNGGL RRSVAPAAPT SCDFSPGDLV WAKMEGYPWW PCLVYNHPFD ...String: MAHHHHHHHH HHGSAASWSH PQFEKGGGSG GGSGGGSWSH PQFEKGALEV LFQGPSRQST LYSFFPKSPA LSDANKASAR ASREGGRAAA APEASPSPGG DAAWSEAGPG PRPLARSASP PKAKNLNGGL RRSVAPAAPT SCDFSPGDLV WAKMEGYPWW PCLVYNHPFD GTFIREKGKS VRVHVQFFDD SPTRGWVSKR LLKPYTGSKS KEAQKGGHFY SAKPEILRAM QRADEALNKD KIKRLELAVC DEPSEPEEEE EMEVGTTYVT DKSEEDNEIE SEEEVQPKTQ GSRRSSRQIK KRRVISDSES DIGGSDVEFK PDTKEEGSSD EISSGVGDSE SEGLNSPVKV ARKRKRMVTG NGSLKRKSSR KETPSATKQA TSISSETKNT LRAFSAPQNS ESQAHVSGGG DDSSRPTVWY HETLEWLKEE KRRDEHRRRP DHPDFDASTL YVPEDFLNSC TPGMRKWWQI KSQNFDLVIC YKVGKFYELY HMDALIGVSE LGLVFMKGNW AHSGFPEIAF GRYSDSLVQK GYKVARVEQT ETPEMMEARC RKMAHISKYD RVVRREICRI ITKGTQTYSV LEGDPSENYS KYLLSLKEKE EDSSGHTRAY GVCFVDTSLG KFFIGQFSDD RHCSRFRTLV AHYPPVQVLF EKGNLSKETK TILKSSLSCS LQEGLIPGSQ FWDASKTLRT LLEEEYFREK LSDGIGVMLP QVLKGMTSES DSIGLTPGEK SELALSALGG CVFYLKKCLI DQELLSMANF EEYIPLDSDT VSTTRSGAIF TKAYQRMVLD AVTLNNLEIF LNGTNGSTEG TLLERVDTCH TPFGKRLLKQ WLCAPLCNHY AINDRLDAIE DLMVVPDKIS EVVELLKKLP DLERLLSKIH NVGSPLKSQN HPDSRAIMYE ETTYSKKKII DFLSALEGFK VMCKIIGIME EVADGFKSKI LKQVISLQTK NPEGRFPDLT VELNRWDTAF DHEKARKTGL ITPKAGFDSD YDQALADIRE NEQSLLEYLE KQRNRIGCRT IVYWGIGRNR YQLEIPENFT TRNLPEEYEL KSTKKGCKRY WTKTIEKKLA NLINAEERRD VSLKDCMRRL FYNFDKNYKD WQSAVECIAV LDVLLCLANY SRGGDGPMCR PVILLPEDTP PFLELKGSRH PCITKTFFGD DFIPNDILIG CEEEEQENGK AYCVLVTGPN MGGKSTLMRQ AGLLAVMAQM GCYVPAEVCR LTPIDRVFTR LGASDRIMSG ESTFFVELSE TASILMHATA HSLVLVDELG RGTATFDGTA IANAVVKELA ETIKCRTLFS THYHSLVEDY SQNVAVRLGH MACMVENECE DPSQETITFL YKFIKGACPK SYGFNAARLA NLPEEVIQKG HRKAREFEKM NQSLRLFREV CLASERSTVD AEAVHKLLTL IKEL |
-Macromolecule #3: DNA (50-mer)
| Macromolecule | Name: DNA (50-mer) / type: dna / ID: 3 / Details: contains a GT mismatch / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Sequence | String: CTTAGCTTAG GATCATCGAG GATCGAGCTC GGTGCAATTC AGCGGTACCC |
-Macromolecule #4: DNA (50-mer)
| Macromolecule | Name: DNA (50-mer) / type: dna / ID: 4 / Details: contains a GT mismatch / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Sequence | String: GGGTACCGCT GAATTGCACC GAGCTTGATC CTCGATGATC CTAAGCTAAG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.023 kPa | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 2 s blotting with force 0. | ||||||||||||||||||
| Details | 1 uM MutSalpha with 2-fold excess mismatched DNA and 1 mM ADP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Details | collected at 200 kV Talos Arctica at Utrecht University, the Netherlands |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 2608 / Average exposure time: 7.8 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 30.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.2 µm / Nominal defocus min: 1.6 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)