+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Single particle structure of Atg18-WT | |||||||||
 Map data Map data | sharp map from CryoSPARC softwarew | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | autophagy / membrane remodeling / PIP binding / PI3P / PI(3 / 5)P2 / lipid binding protein / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of phosphatidylinositol biosynthetic process / PAS complex / 1-phosphatidyl-1D-myo-inositol 3,5-bisphosphate metabolic process / phagophore / positive regulation of vacuole organization / vacuolar protein processing / glycophagy / Macroautophagy / cytoplasm to vacuole targeting by the Cvt pathway / nucleophagy ...regulation of phosphatidylinositol biosynthetic process / PAS complex / 1-phosphatidyl-1D-myo-inositol 3,5-bisphosphate metabolic process / phagophore / positive regulation of vacuole organization / vacuolar protein processing / glycophagy / Macroautophagy / cytoplasm to vacuole targeting by the Cvt pathway / nucleophagy / pexophagy / protein localization to phagophore assembly site / phagophore assembly site membrane / late endosome to vacuole transport / piecemeal microautophagy of the nucleus / phosphatidylinositol-3-phosphate binding / fungal-type vacuole membrane / phagophore assembly site / phosphatidylinositol-4-phosphate binding / phosphatidylinositol-3,5-bisphosphate binding / vacuolar membrane / extrinsic component of membrane / autophagosome assembly / ubiquitin binding / cell periphery / macroautophagy / endosome membrane / endosome / protein-containing complex / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
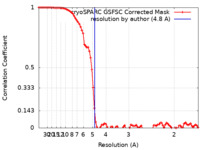
| Method | single particle reconstruction / cryo EM / Resolution: 4.8 Å | |||||||||
 Authors Authors | Mann D / Fromm S / Martinez-Sanchez A / Gopaldass N / Mayer A / Sachse C | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structures of Atg18 oligomers reveal a tilted structural scaffold for Atg2 at the isolation membrane Authors: Mann D / Fromm SA / Martinez-Sanchez A / Gopaldass N / Mayer A / Sachse C | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15411.map.gz emd_15411.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15411-v30.xml emd-15411-v30.xml emd-15411.xml emd-15411.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15411_fsc.xml emd_15411_fsc.xml | 9.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_15411.png emd_15411.png | 86.1 KB | ||
| Masks |  emd_15411_msk_1.map emd_15411_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15411.cif.gz emd-15411.cif.gz | 5.6 KB | ||
| Others |  emd_15411_half_map_1.map.gz emd_15411_half_map_1.map.gz emd_15411_half_map_2.map.gz emd_15411_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15411 http://ftp.pdbj.org/pub/emdb/structures/EMD-15411 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15411 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15411 | HTTPS FTP |
-Validation report
| Summary document |  emd_15411_validation.pdf.gz emd_15411_validation.pdf.gz | 844.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15411_full_validation.pdf.gz emd_15411_full_validation.pdf.gz | 844.2 KB | Display | |
| Data in XML |  emd_15411_validation.xml.gz emd_15411_validation.xml.gz | 16.2 KB | Display | |
| Data in CIF |  emd_15411_validation.cif.gz emd_15411_validation.cif.gz | 21 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15411 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15411 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15411 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15411 | HTTPS FTP |
-Related structure data
| Related structure data |  8afxMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15411.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15411.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharp map from CryoSPARC softwarew | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8389 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15411_msk_1.map emd_15411_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_15411_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_15411_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Atg18
| Entire | Name: Atg18 |
|---|---|
| Components |
|
-Supramolecule #1: Atg18
| Supramolecule | Name: Atg18 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Autophagy-related protein 18
| Macromolecule | Name: Autophagy-related protein 18 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 54.62241 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SPTINFINFN QTGTCISLGT SKGFKIFNCE PFGKFYSEDS GGYAIVEMLF STSLLALVGI GDQPALSPRR LRIINTKKHS IICEVTFPT SILSVKMNKS RLVVLLQEQI YIYDINTMRL LHTIETNPNP RGLMAMSPSV ANSYLVYPSP PKVINSEIKA H ATTNNITL ...String: SPTINFINFN QTGTCISLGT SKGFKIFNCE PFGKFYSEDS GGYAIVEMLF STSLLALVGI GDQPALSPRR LRIINTKKHS IICEVTFPT SILSVKMNKS RLVVLLQEQI YIYDINTMRL LHTIETNPNP RGLMAMSPSV ANSYLVYPSP PKVINSEIKA H ATTNNITL SVGGNTETSF KRDQQDAGHS DISDLDQYSS FTKRDDADPT SSNGGNSSII KNGDVIVFNL ETLQPTMVIE AH KGEIAAM AISFDGTLMA TASDKGTIIR VFDIETGDKI YQFRRGTYAT RIYSISFSED SQYLAVTGSS KTVHIFKLGH SMS NNKLDS DDSNMEEAAA DDSSLDTTSI DALSDEENPT RLAREPYVDA SRKTMGRMIR YSSQKLSRRA ARTLGQIFPI KVTS LLESS RHFASLKLPV ETNSHVMTIS SIGSPIDIDT SEYPELFETG NSASTESYHE PVMKMVPIRV VSSDGYLYNF VMDPE RGGD CLILSQYSIL M UniProtKB: Autophagy-related protein 18 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS TALOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: COUNTING / Number real images: 5117 / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)