+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
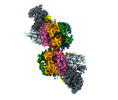
| Title | S. cerevisiae CMGE dimer nucleating origin DNA melting | |||||||||||||||||||||||||||||||||
 Map data Map data | Consensus map of CMGE dimer model generated from EMD EMD-13978 | |||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||
 Keywords Keywords | DNA replication / helicase / initiation / DNA origin / REPLICATION | |||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology information: / cell cycle / DNA-templated DNA replication maintenance of fidelity / gene conversion / Unwinding of DNA / DNA replication initiation / replication fork protection complex / epsilon DNA polymerase complex / MCM core complex / Assembly of the pre-replicative complex ...: / cell cycle / DNA-templated DNA replication maintenance of fidelity / gene conversion / Unwinding of DNA / DNA replication initiation / replication fork protection complex / epsilon DNA polymerase complex / MCM core complex / Assembly of the pre-replicative complex / Switching of origins to a post-replicative state / DNA strand elongation involved in mitotic DNA replication / GINS complex / MCM complex binding / mitotic DNA replication preinitiation complex assembly / nuclear DNA replication / premeiotic DNA replication / pre-replicative complex assembly involved in nuclear cell cycle DNA replication / nucleotide-excision repair, DNA gap filling / Activation of the pre-replicative complex / SUMO binding / mitotic DNA replication / CMG complex / nuclear pre-replicative complex / DNA replication proofreading / : / DNA replication preinitiation complex / Activation of ATR in response to replication stress / single-stranded DNA 3'-5' DNA exonuclease activity / mitotic DNA replication checkpoint signaling / double-strand break repair via break-induced replication / MCM complex / mitotic DNA replication initiation / mitotic intra-S DNA damage checkpoint signaling / silent mating-type cassette heterochromatin formation / single-stranded DNA helicase activity / mitotic sister chromatid cohesion / Hydrolases; Acting on ester bonds; Exodeoxyribonucleases producing 5'-phosphomonoesters / DNA strand elongation involved in DNA replication / leading strand elongation / nuclear replication fork / 3'-5' DNA helicase activity / Dual incision in TC-NER / DNA replication origin binding / DNA replication initiation / error-prone translesion synthesis / subtelomeric heterochromatin formation / base-excision repair, gap-filling / DNA helicase activity / replication fork / helicase activity / base-excision repair / DNA-templated DNA replication / double-strand break repair via nonhomologous end joining / peroxisome / double-strand break repair / mitotic cell cycle / single-stranded DNA binding / 4 iron, 4 sulfur cluster binding / double-stranded DNA binding / DNA-directed DNA polymerase / DNA helicase / DNA-directed DNA polymerase activity / chromosome, telomeric region / DNA replication / nucleotide binding / hydrolase activity / mRNA binding / chromatin binding / ATP hydrolysis activity / DNA binding / zinc ion binding / nucleoplasm / ATP binding / metal ion binding / nucleus / cytoplasm Similarity search - Function | |||||||||||||||||||||||||||||||||
| Biological species |  | |||||||||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||||||||||||||||||||||||||
 Authors Authors | Lewis JS / Sousa JS | |||||||||||||||||||||||||||||||||
| Funding support | European Union,  France, France,  United Kingdom, 10 items United Kingdom, 10 items
| |||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Mechanism of replication origin melting nucleated by CMG helicase assembly. Authors: Jacob S Lewis / Marta H Gross / Joana Sousa / Sarah S Henrikus / Julia F Greiwe / Andrea Nans / John F X Diffley / Alessandro Costa /  Abstract: The activation of eukaryotic origins of replication occurs in temporally separated steps to ensure that chromosomes are copied only once per cell cycle. First, the MCM helicase is loaded onto duplex ...The activation of eukaryotic origins of replication occurs in temporally separated steps to ensure that chromosomes are copied only once per cell cycle. First, the MCM helicase is loaded onto duplex DNA as an inactive double hexamer. Activation occurs after the recruitment of a set of firing factors that assemble two Cdc45-MCM-GINS (CMG) holo-helicases. CMG formation leads to the underwinding of DNA on the path to the establishment of the replication fork, but whether DNA becomes melted at this stage is unknown. Here we use cryo-electron microscopy to image ATP-dependent CMG assembly on a chromatinized origin, reconstituted in vitro with purified yeast proteins. We find that CMG formation disrupts the double hexamer interface and thereby exposes duplex DNA in between the two CMGs. The two helicases remain tethered, which gives rise to a splayed dimer, with implications for origin activation and replisome integrity. Inside each MCM ring, the double helix becomes untwisted and base pairing is broken. This comes as the result of ATP-triggered conformational changes in MCM that involve DNA stretching and protein-mediated stabilization of three orphan bases. Mcm2 pore-loop residues that engage DNA in our structure are dispensable for double hexamer loading and CMG formation, but are essential to untwist the DNA and promote replication. Our results explain how ATP binding nucleates origin DNA melting by the CMG and maintains replisome stability at initiation. | |||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14439.map.gz emd_14439.map.gz | 43.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14439-v30.xml emd-14439-v30.xml emd-14439.xml emd-14439.xml | 40.2 KB 40.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14439.png emd_14439.png | 83.3 KB | ||
| Filedesc metadata |  emd-14439.cif.gz emd-14439.cif.gz | 13.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14439 http://ftp.pdbj.org/pub/emdb/structures/EMD-14439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14439 | HTTPS FTP |
-Related structure data
| Related structure data |  7z13MC  7qhsC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14439.map.gz / Format: CCP4 / Size: 60.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14439.map.gz / Format: CCP4 / Size: 60.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Consensus map of CMGE dimer model generated from EMD EMD-13978 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : S. cerevisiae CMGE dimer nucleating origin DNA melting
+Supramolecule #1: S. cerevisiae CMGE dimer nucleating origin DNA melting
+Macromolecule #1: DNA replication licensing factor MCM2
+Macromolecule #2: DNA replication licensing factor MCM3
+Macromolecule #3: DNA replication licensing factor MCM4
+Macromolecule #4: DNA helicase
+Macromolecule #5: DNA replication licensing factor MCM6
+Macromolecule #6: DNA replication licensing factor MCM7
+Macromolecule #9: DNA replication complex GINS protein PSF3
+Macromolecule #10: DNA replication complex GINS protein SLD5
+Macromolecule #11: Cell division control protein 45
+Macromolecule #12: DNA polymerase epsilon subunit B
+Macromolecule #13: DNA replication complex GINS protein PSF1
+Macromolecule #14: DNA replication complex GINS protein PSF2
+Macromolecule #15: DNA polymerase epsilon catalytic subunit A
+Macromolecule #7: DNA (53-MER)
+Macromolecule #8: DNA (53-MER)
+Macromolecule #16: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #17: ZINC ION
+Macromolecule #18: MAGNESIUM ION
+Macromolecule #19: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
| Details | four microlitres of sample was applied on a grid and incubated for 2 min at room temperature before blotting with filter paper for 5.5 s and plunge-freezing in liquid ethane. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 65286 / Average exposure time: 10.0 sec. / Average electron dose: 1.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.4 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.4 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 71348 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | One additional base pair has been built to connect the DNA molecules from the two individual symmetry expanded monomers. |
| Refinement | Protocol: RIGID BODY FIT |
| Output model |  PDB-7z13: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)