[English] 日本語
 Yorodumi
Yorodumi- EMDB-14289: Bovine complex I in the presence of IM1761092, deactive class iv ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bovine complex I in the presence of IM1761092, deactive class iv (Hydrophilic domain) | |||||||||
 Map data Map data | Hydrophilic domain globally sharpened | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
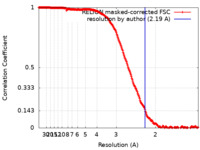
| Method | single particle reconstruction / cryo EM / Resolution: 2.19 Å | |||||||||
 Authors Authors | Bridges HR / Blaza JN / Yin Z / Chung I / Hirst J | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Structural basis of mammalian respiratory complex I inhibition by medicinal biguanides. Authors: Hannah R Bridges / James N Blaza / Zhan Yin / Injae Chung / Michael N Pollak / Judy Hirst /   Abstract: The molecular mode of action of biguanides, including the drug metformin, which is widely used in the treatment of diabetes, is incompletely characterized. Here, we define the inhibitory drug-target ...The molecular mode of action of biguanides, including the drug metformin, which is widely used in the treatment of diabetes, is incompletely characterized. Here, we define the inhibitory drug-target interaction(s) of a model biguanide with mammalian respiratory complex I by combining cryo-electron microscopy and enzyme kinetics. We interpret these data to explain the selectivity of biguanide binding to different enzyme states. The primary inhibitory site is in an amphipathic region of the quinone-binding channel, and an additional binding site is in a pocket on the intermembrane-space side of the enzyme. An independent local chaotropic interaction, not previously described for any drug, displaces a portion of a key helix in the membrane domain. Our data provide a structural basis for biguanide action and enable the rational design of medicinal biguanides. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14289.map.gz emd_14289.map.gz | 1003.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14289-v30.xml emd-14289-v30.xml emd-14289.xml emd-14289.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14289_fsc.xml emd_14289_fsc.xml | 23.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_14289.png emd_14289.png | 79.6 KB | ||
| Masks |  emd_14289_msk_1.map emd_14289_msk_1.map | 1.1 GB |  Mask map Mask map | |
| Others |  emd_14289_half_map_1.map.gz emd_14289_half_map_1.map.gz emd_14289_half_map_2.map.gz emd_14289_half_map_2.map.gz | 891.8 MB 891.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14289 http://ftp.pdbj.org/pub/emdb/structures/EMD-14289 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14289 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14289 | HTTPS FTP |
-Validation report
| Summary document |  emd_14289_validation.pdf.gz emd_14289_validation.pdf.gz | 881.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14289_full_validation.pdf.gz emd_14289_full_validation.pdf.gz | 881.4 KB | Display | |
| Data in XML |  emd_14289_validation.xml.gz emd_14289_validation.xml.gz | 30.8 KB | Display | |
| Data in CIF |  emd_14289_validation.cif.gz emd_14289_validation.cif.gz | 41.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14289 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14289 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14289 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14289 | HTTPS FTP |
-Related structure data
| Related structure data |  7qsdC  7r41C  7r42C  7r43C  7r44C  7r45C  7r46C  7r47C  7r48C  7r4cC  7r4dC  7r4fC  7r4gC C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14289.map.gz / Format: CCP4 / Size: 1.1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14289.map.gz / Format: CCP4 / Size: 1.1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Hydrophilic domain globally sharpened | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.731 Å | ||||||||||||||||||||||||||||||||||||
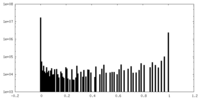
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14289_msk_1.map emd_14289_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
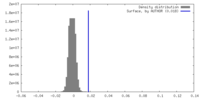
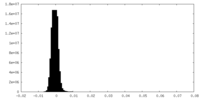
| Density Histograms |
-Half map: Hydrophilic domain halfmap1
| File | emd_14289_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Hydrophilic domain halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
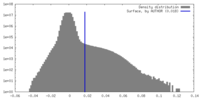
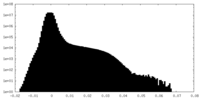
| Density Histograms |
-Half map: Hydrophilic domain halfmap2
| File | emd_14289_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Hydrophilic domain halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
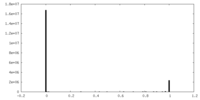
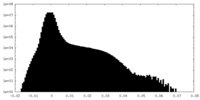
| Density Histograms |
- Sample components
Sample components
-Entire : NADH Ubiquinone oxidoreductase (Complex I)
| Entire | Name: NADH Ubiquinone oxidoreductase (Complex I) |
|---|---|
| Components |
|
-Supramolecule #1: NADH Ubiquinone oxidoreductase (Complex I)
| Supramolecule | Name: NADH Ubiquinone oxidoreductase (Complex I) / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#45 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.14 Component:
| ||||||||||||
| Grid | Model: UltrAuFoil R0.6/1 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. / Pretreatment - Atmosphere: AIR Details: Grid also covalently modified by peg-thiol for 48 hours in a nitrogen atmosphere. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)