+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Nucleosome 3 of the 4x197 nucleosome array containing H1 | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Chromatin / Nucleosomes / Linker Histone / DNA BINDING PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / telomere organization ...negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / telomere organization / Interleukin-7 signaling / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / RNA Polymerase I Promoter Opening / Inhibition of DNA recombination at telomere / Assembly of the ORC complex at the origin of replication / Meiotic synapsis / SUMOylation of chromatin organization proteins / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / innate immune response in mucosa / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / HDMs demethylate histones / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / Negative Regulation of CDH1 Gene Transcription / NoRC negatively regulates rRNA expression / PKMTs methylate histone lysines / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / Pre-NOTCH Transcription and Translation / Meiotic recombination / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Transcriptional regulation of granulopoiesis / Metalloprotease DUBs / RMTs methylate histone arginines / HCMV Early Events / structural constituent of chromatin / UCH proteinases / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / heterochromatin formation / nucleosome assembly / antibacterial humoral response / E3 ubiquitin ligases ubiquitinate target proteins / HATs acetylate histones / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / Factors involved in megakaryocyte development and platelet production / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / chromatin organization / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Processing of DNA double-strand break ends / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / killing of cells of another organism / gene expression / defense response to Gram-negative bacterium / chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / chromatin binding / negative regulation of transcription by RNA polymerase II / protein-containing complex / : / DNA binding / RNA binding / extracellular exosome / extracellular region / nucleoplasm / membrane / nucleus / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / synthetic construct (人工物) Homo sapiens (ヒト) / synthetic construct (人工物) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.3 Å | |||||||||
 データ登録者 データ登録者 | Dombrowski M / Cramer P | |||||||||
| 資金援助 | European Union, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nat Struct Mol Biol / 年: 2022 ジャーナル: Nat Struct Mol Biol / 年: 2022タイトル: Histone H1 binding to nucleosome arrays depends on linker DNA length and trajectory. 著者: Marco Dombrowski / Maik Engeholm / Christian Dienemann / Svetlana Dodonova / Patrick Cramer /  要旨: Throughout the genome, nucleosomes often form regular arrays that differ in nucleosome repeat length (NRL), occupancy of linker histone H1 and transcriptional activity. Here, we report cryo-EM ...Throughout the genome, nucleosomes often form regular arrays that differ in nucleosome repeat length (NRL), occupancy of linker histone H1 and transcriptional activity. Here, we report cryo-EM structures of human H1-containing tetranucleosome arrays with four physiologically relevant NRLs. The structures show a zig-zag arrangement of nucleosomes, with nucleosomes 1 and 3 forming a stack. H1 binding to stacked nucleosomes depends on the NRL, whereas H1 always binds to the non-stacked nucleosomes 2 and 4. Short NRLs lead to altered trajectories of linker DNA, and these altered trajectories sterically impair H1 binding to the stacked nucleosomes in our structures. As the NRL increases, linker DNA trajectories relax, enabling H1 contacts and binding. Our results provide an explanation for why arrays with short NRLs are depleted of H1 and suited for transcription, whereas arrays with long NRLs show full H1 occupancy and can form transcriptionally silent heterochromatin regions. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_13374.map.gz emd_13374.map.gz | 62.1 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-13374-v30.xml emd-13374-v30.xml emd-13374.xml emd-13374.xml | 15.2 KB 15.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_13374_fsc.xml emd_13374_fsc.xml | 10.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_13374.png emd_13374.png | 45.3 KB | ||
| Filedesc metadata |  emd-13374.cif.gz emd-13374.cif.gz | 5.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13374 http://ftp.pdbj.org/pub/emdb/structures/EMD-13374 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13374 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13374 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7pffMC  7petC  7peuC  7pevC  7pewC  7pexC  7peyC  7pezC  7pf0C  7pf2C  7pf3C  7pf4C  7pf5C  7pf6C  7pfaC  7pfcC  7pfdC  7pfeC  7pftC  7pfuC  7pfvC  7pfwC  7pfxC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_13374.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_13374.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
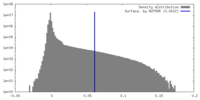
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : Nucleosome 3 from 4x197 nucleosome array
| 全体 | 名称: Nucleosome 3 from 4x197 nucleosome array |
|---|---|
| 要素 |
|
-超分子 #1: Nucleosome 3 from 4x197 nucleosome array
| 超分子 | 名称: Nucleosome 3 from 4x197 nucleosome array / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|
-分子 #1: Histone H3.2
| 分子 | 名称: Histone H3.2 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 15.389036 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MARTKQTARK STGGKAPRKQ LATKAARKSA PATGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSSA VMALQEASEA YLVGLFEDTN LAAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.2 |
-分子 #2: Histone H4
| 分子 | 名称: Histone H4 / タイプ: protein_or_peptide / ID: 2 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 11.394426 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-分子 #3: Histone H2A type 1-B/E
| 分子 | 名称: Histone H2A type 1-B/E / タイプ: protein_or_peptide / ID: 3 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 16.344873 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: HHHHHHENLY FQSNAPWMSG RGKQGGKARA KAKTRSSRAG LQFPVGRVHR LLRKGNYSER VGAGAPVYLA AVLEYLTAEI LELAGNAAR DNKKTRIIPR HLQLAIRNDE ELNKLLGRVT IAQGGVLPNI QAVLLPKKTE SHHKAKGK UniProtKB: Histone H2A type 1-B/E |
-分子 #4: Histone H2B type 1-K
| 分子 | 名称: Histone H2B type 1-K / タイプ: protein_or_peptide / ID: 4 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 13.921213 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSV YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSAK UniProtKB: Histone H2B type 1-K |
-分子 #5: DNA (167-MER)
| 分子 | 名称: DNA (167-MER) / タイプ: dna / ID: 5 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種: synthetic construct (人工物) |
| 分子量 | 理論値: 51.333703 KDa |
| 配列 | 文字列: (DG)(DG)(DC)(DC)(DG)(DC)(DC)(DA)(DT)(DA) (DC)(DT)(DG)(DG)(DA)(DG)(DA)(DA)(DT)(DC) (DC)(DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG) (DA)(DG)(DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA) (DA) (DT)(DT)(DG)(DG)(DT) ...文字列: (DG)(DG)(DC)(DC)(DG)(DC)(DC)(DA)(DT)(DA) (DC)(DT)(DG)(DG)(DA)(DG)(DA)(DA)(DT)(DC) (DC)(DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG) (DA)(DG)(DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA) (DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT) (DA)(DG)(DA)(DC)(DA)(DG)(DC)(DT)(DC)(DT) (DA)(DG) (DC)(DA)(DC)(DC)(DG)(DC)(DT) (DT)(DA)(DA)(DA)(DC)(DG)(DC)(DA)(DC)(DG) (DT)(DA)(DC) (DG)(DC)(DG)(DC)(DT)(DG) (DT)(DC)(DC)(DC)(DC)(DC)(DG)(DC)(DG)(DT) (DT)(DT)(DT)(DA) (DA)(DC)(DC)(DG)(DC) (DC)(DA)(DA)(DG)(DG)(DG)(DG)(DA)(DT)(DT) (DA)(DC)(DT)(DC)(DC) (DC)(DT)(DA)(DG) (DT)(DC)(DT)(DC)(DC)(DA)(DG)(DG)(DC)(DA) (DC)(DG)(DT)(DG)(DT)(DC) (DA)(DG)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DC)(DA)(DT)(DC) (DC)(DT)(DG)(DT)(DC)(DA)(DT) (DG)(DT) (DA)(DA)(DG)(DT)(DA) |
-分子 #6: DNA (167-MER)
| 分子 | 名称: DNA (167-MER) / タイプ: dna / ID: 6 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種: synthetic construct (人工物) |
| 分子量 | 理論値: 51.773973 KDa |
| 配列 | 文字列: (DT)(DA)(DC)(DT)(DT)(DA)(DC)(DA)(DT)(DG) (DA)(DC)(DA)(DG)(DG)(DA)(DT)(DG)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA) (DC)(DA)(DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG) (DG) (DA)(DG)(DA)(DC)(DT) ...文字列: (DT)(DA)(DC)(DT)(DT)(DA)(DC)(DA)(DT)(DG) (DA)(DC)(DA)(DG)(DG)(DA)(DT)(DG)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA) (DC)(DA)(DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG) (DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG) (DG)(DA)(DG)(DT)(DA)(DA)(DT)(DC)(DC)(DC) (DC)(DT) (DT)(DG)(DG)(DC)(DG)(DG)(DT) (DT)(DA)(DA)(DA)(DA)(DC)(DG)(DC)(DG)(DG) (DG)(DG)(DG) (DA)(DC)(DA)(DG)(DC)(DG) (DC)(DG)(DT)(DA)(DC)(DG)(DT)(DG)(DC)(DG) (DT)(DT)(DT)(DA) (DA)(DG)(DC)(DG)(DG) (DT)(DG)(DC)(DT)(DA)(DG)(DA)(DG)(DC)(DT) (DG)(DT)(DC)(DT)(DA) (DC)(DG)(DA)(DC) (DC)(DA)(DA)(DT)(DT)(DG)(DA)(DG)(DC)(DG) (DG)(DC)(DC)(DT)(DC)(DG) (DG)(DC)(DA) (DC)(DC)(DG)(DG)(DG)(DA)(DT)(DT)(DC)(DT) (DC)(DC)(DA)(DG)(DT)(DA)(DT) (DG)(DG) (DC)(DG)(DG)(DC)(DC) |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.1 mg/mL |
|---|---|
| 緩衝液 | pH: 7 |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 60.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー


































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)