[English] 日本語
 Yorodumi
Yorodumi- EMDB-13036: Cryo-EM structure of the human TRPA1 ion channel in complex with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human TRPA1 ion channel in complex with the antagonist 3-60, conformation 2 | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Transient receptor potential cation channel subfamily A member 1 / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of blood circulation / positive regulation of monoatomic anion transport / cellular response to food / temperature-gated cation channel activity / regulation of neuronal action potential / cellular response to carbon dioxide / osmolarity-sensing monoatomic cation channel activity / stereocilium bundle / detection of chemical stimulus involved in sensory perception of pain / thermoception ...regulation of blood circulation / positive regulation of monoatomic anion transport / cellular response to food / temperature-gated cation channel activity / regulation of neuronal action potential / cellular response to carbon dioxide / osmolarity-sensing monoatomic cation channel activity / stereocilium bundle / detection of chemical stimulus involved in sensory perception of pain / thermoception / urinary bladder smooth muscle contraction / cellular response to toxic substance / cellular response to caffeine / response to pain / TRP channels / calcium ion transmembrane import into cytosol / cellular response to cold / intracellularly gated calcium channel activity / detection of mechanical stimulus involved in sensory perception of pain / voltage-gated calcium channel activity / monoatomic ion transport / sensory perception of pain / response to cold / positive regulation of insulin secretion involved in cellular response to glucose stimulus / calcium ion transmembrane transport / calcium channel activity / cellular response to hydrogen peroxide / intracellular calcium ion homeostasis / cellular response to heat / channel activity / protein homotetramerization / cell surface receptor signaling pathway / apical plasma membrane / response to xenobiotic stimulus / axon / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
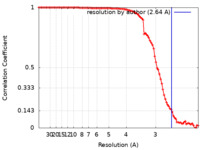
| Method | single particle reconstruction / cryo EM / Resolution: 2.64 Å | |||||||||
 Authors Authors | Grieben M / Pike ACW | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of the human TRPA1 ion channel in complex with the antagonist 3-60 Authors: Grieben M / Saward BG / Pike ACW / Schofield CJ / Carpenter EP | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13036.map.gz emd_13036.map.gz | 41.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13036-v30.xml emd-13036-v30.xml emd-13036.xml emd-13036.xml | 17.6 KB 17.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13036_fsc.xml emd_13036_fsc.xml | 9.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_13036.png emd_13036.png | 121 KB | ||
| Masks |  emd_13036_msk_1.map emd_13036_msk_1.map | 80.2 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13036.cif.gz emd-13036.cif.gz | 6.4 KB | ||
| Others |  emd_13036_half_map_1.map.gz emd_13036_half_map_1.map.gz emd_13036_half_map_2.map.gz emd_13036_half_map_2.map.gz | 74 MB 74 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13036 http://ftp.pdbj.org/pub/emdb/structures/EMD-13036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13036 | HTTPS FTP |
-Related structure data
| Related structure data |  7or0MC  7or1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13036.map.gz / Format: CCP4 / Size: 80.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13036.map.gz / Format: CCP4 / Size: 80.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_13036_msk_1.map emd_13036_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half A
| File | emd_13036_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half B
| File | emd_13036_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Transient receptor potential cation channel subfamily A member 1
| Entire | Name: Transient receptor potential cation channel subfamily A member 1 |
|---|---|
| Components |
|
-Supramolecule #1: Transient receptor potential cation channel subfamily A member 1
| Supramolecule | Name: Transient receptor potential cation channel subfamily A member 1 type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Transient receptor potential cation channel subfamily A member 1
| Macromolecule | Name: Transient receptor potential cation channel subfamily A member 1 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 128.455336 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKRSLRKMWR PGEKKEPQGV VYEDVPDDTE DFKESLKVVF EGSAYGLQNF NKQKKLKTCD DMDTFFLHYA AAEGQIELME KITRDSSLE VLHEMDDYGN TPLHCAVEKN QIESVKFLLS RGANPNLRNF NMMAPLHIAV QGMNNEVMKV LLEHRTIDVN L EGENGNTA ...String: MKRSLRKMWR PGEKKEPQGV VYEDVPDDTE DFKESLKVVF EGSAYGLQNF NKQKKLKTCD DMDTFFLHYA AAEGQIELME KITRDSSLE VLHEMDDYGN TPLHCAVEKN QIESVKFLLS RGANPNLRNF NMMAPLHIAV QGMNNEVMKV LLEHRTIDVN L EGENGNTA VIIACTTNNS EALQILLNKG AKPCKSNKWG CFPIHQAAFS GSKECMEIIL RFGEEHGYSR QLHINFMNNG KA TPLHLAV QNGDLEMIKM CLDNGAQIDP VEKGRCTAIH FAATQGATEI VKLMISSYSG SVDIVNTTDG CHETMLHRAS LFD HHELAD YLISVGADIN KIDSEGRSPL ILATASASWN IVNLLLSKGA QVDIKDNFGR NFLHLTVQQP YGLKNLRPEF MQMQ QIKEL VMDEDNDGCT PLHYACRQGG PGSVNNLLGF NVSIHSKSKD KKSPLHFAAS YGRINTCQRL LQDISDTRLL NEGDL HGMT PLHLAAKNGH DKVVQLLLKK GALFLSDHNG WTALHHASMG GYTQTMKVIL DTNLKCTDRL DEDGNTALHF AAREGH AKA VALLLSHNAD IVLNKQQASF LHLALHNKRK EVVLTIIRSK RWDECLKIFS HNSPGNKCPI TEMIEYLPEC MKVLLDF CM LHSTEDKSCR DYYIEYNFKY LQCPLEFTKK TPTQDVIYEP LTALNAMVQN NRIELLNHPV CKEYLLMKWL AYGFRAHM M NLGSYCLGLI PMTILVVNIK PGMAFNSTGI INETSDHSEI LDTTNSYLIK TCMILVFLSS IFGYCKEAGQ IFQQKRNYF MDISNVLEWI IYTTGIIFVL PLFVEIPAHL QWQCGAIAVY FYWMNFLLYL QRFENCGIFI VMLEVILKTL LRSTVVFIFL LLAFGLSFY ILLNLQDPFS SPLLSIIQTF SMMLGDINYR ESFLEPYLRN ELAHPVLSFA QLVSFTIFVP IVLMNLLIGL A VGDIAEVQ KHASLKRIAM QVELHTSLEK KLPLWFLRKV DQKSTIVYPN KPRSGGMLFH IFCFLFCTGE IRQEIPNADK SL EMEILKQ KYRLKDLTFL LEKQHELIKL IIQKMEIISE TEDDDSHCSF QDRFKKEQME QRNSRWNTVL RAVKAKTHHL EPA ENLYFQ UniProtKB: Transient receptor potential cation channel subfamily A member 1 |
-Macromolecule #3: ~{N}-[4-[4-[(azanylidene-$l^{4}-azanylidene)amino]phenyl]-1,3-thi...
| Macromolecule | Name: ~{N}-[4-[4-[(azanylidene-$l^{4}-azanylidene)amino]phenyl]-1,3-thiazol-2-yl]-2-[1,3-dimethyl-2,6-bis(oxidanylidene)purin-7-yl]ethanamide type: ligand / ID: 3 / Number of copies: 4 / Formula: 0IG |
|---|---|
| Molecular weight | Theoretical: 438.443 Da |
| Chemical component information |  ChemComp-0IG: |
-Macromolecule #4: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 4 / Number of copies: 8 / Formula: PC1 |
|---|---|
| Molecular weight | Theoretical: 790.145 Da |
| Chemical component information |  ChemComp-PC1: |
-Macromolecule #5: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 5 / Number of copies: 5 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #7: INOSITOL HEXAKISPHOSPHATE
| Macromolecule | Name: INOSITOL HEXAKISPHOSPHATE / type: ligand / ID: 7 / Number of copies: 4 / Formula: IHP |
|---|---|
| Molecular weight | Theoretical: 660.035 Da |
| Chemical component information |  ChemComp-IHP: |
-Macromolecule #8: water
| Macromolecule | Name: water / type: ligand / ID: 8 / Number of copies: 52 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Average electron dose: 45.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 1.4000000000000001 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)