[English] 日本語
 Yorodumi
Yorodumi- EMDB-1216: Cryo-electron tomographic structure of an immunodeficiency virus ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1216 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron tomographic structure of an immunodeficiency virus envelope complex in situ. | |||||||||
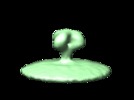
 Map data Map data | This is the volume of the averaged SIV envelope complex | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Simian immunodeficiency virus / viral envelope (unknown) Simian immunodeficiency virus / viral envelope (unknown) | |||||||||
| Method | subtomogram averaging / cryo EM / negative staining / Resolution: 28.0 Å | |||||||||
 Authors Authors | Zanetti G / Briggs JAG / Gruenewald K / Sattentau Q / Fuller SD | |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2006 Journal: PLoS Pathog / Year: 2006Title: Cryo-electron tomographic structure of an immunodeficiency virus envelope complex in situ. Authors: Giulia Zanetti / John A G Briggs / Kay Grünewald / Quentin J Sattentau / Stephen D Fuller /  Abstract: The envelope glycoprotein (Env) complexes of the human and simian immunodeficiency viruses (HIV and SIV, respectively) mediate viral entry and are a target for neutralizing antibodies. The receptor ...The envelope glycoprotein (Env) complexes of the human and simian immunodeficiency viruses (HIV and SIV, respectively) mediate viral entry and are a target for neutralizing antibodies. The receptor binding surfaces of Env are in large part sterically occluded or conformationally masked prior to receptor binding. Knowledge of the unliganded, trimeric Env structure is key for an understanding of viral entry and immune escape, and for the design of vaccines to elicit neutralizing antibodies. We have used cryo-electron tomography and averaging to obtain the structure of the SIV Env complex prior to fusion. Our result reveals novel details of Env organisation, including tight interaction between monomers in the gp41 trimer, associated with a three-lobed, membrane-distal gp120 trimer. A cavity exists at the gp41-gp120 trimer interface. Our model for the spike structure agrees with previously predicted interactions between gp41 monomers, and furthers our understanding of gp120 interactions within an intact spike. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1216.map.gz emd_1216.map.gz | 519.4 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1216-v30.xml emd-1216-v30.xml emd-1216.xml emd-1216.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| Images |  1216.gif 1216.gif | 7.4 KB | ||
| Masks |  emd_1216_msk_1.map emd_1216_msk_1.map emd_1216_msk_2.map emd_1216_msk_2.map | 977.7 KB 977.7 KB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1216 http://ftp.pdbj.org/pub/emdb/structures/EMD-1216 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1216 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1216 | HTTPS FTP |
-Validation report
| Summary document |  emd_1216_validation.pdf.gz emd_1216_validation.pdf.gz | 212.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1216_full_validation.pdf.gz emd_1216_full_validation.pdf.gz | 211.4 KB | Display | |
| Data in XML |  emd_1216_validation.xml.gz emd_1216_validation.xml.gz | 5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1216 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1216 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1216 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1216 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1216.map.gz / Format: CCP4 / Size: 955.1 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1216.map.gz / Format: CCP4 / Size: 955.1 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is the volume of the averaged SIV envelope complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.47 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
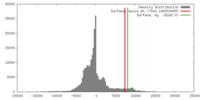
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Segmentation: The mask for the averaged SIV envelope complex
| Annotation | The mask for the averaged SIV envelope complex | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_1216_msk_1.map emd_1216_msk_1.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
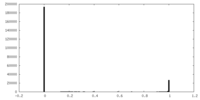
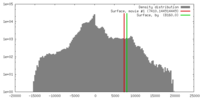
| Density Histograms |
-Segmentation: Protein with the mask applied is the simple...
| Annotation | Protein with the mask applied is the simple multiplication of the spike protein map (emd_1216.map) with the mask (emd_1216_mask_1.map) | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_1216_msk_2.map emd_1216_msk_2.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
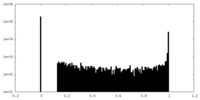
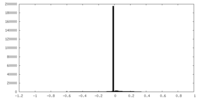
| Density Histograms |
- Sample components
Sample components
-Entire : Simian Immunodeficiency Virus mne Envelope Complex
| Entire | Name: Simian Immunodeficiency Virus mne Envelope Complex |
|---|---|
| Components |
|
-Supramolecule #1000: Simian Immunodeficiency Virus mne Envelope Complex
| Supramolecule | Name: Simian Immunodeficiency Virus mne Envelope Complex / type: sample / ID: 1000 Details: The envelope complex wasn't purified but it was analysed on the virus Oligomeric state: trimer of Gp120 and trimer of Gp41 and membrane Number unique components: 3 |
|---|---|
| Molecular weight | Experimental: 500 KDa / Theoretical: 500 KDa Method: sum of individual components is an underestimate due to the presence of the membrane |
-Supramolecule #1: viral envelope
| Supramolecule | Name: viral envelope / type: virus / ID: 1 / Name.synonym: viral membrane / Details: viral membrane / Sci species name: viral envelope / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: viral membrane |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Name: membrane / Diameter: 1200 Å |
-Macromolecule #1: gp120
| Macromolecule | Name: gp120 / type: protein_or_peptide / ID: 1 / Name.synonym: SIV envelope receptor binding protein / Details: trimeric / Number of copies: 3 / Oligomeric state: trimer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Simian immunodeficiency virus / Strain: SIVMneCL8 / synonym: SIVmne / Cell: HuT 78 / Location in cell: viral membrane Simian immunodeficiency virus / Strain: SIVMneCL8 / synonym: SIVmne / Cell: HuT 78 / Location in cell: viral membrane |
| Molecular weight | Experimental: 120 KDa / Theoretical: 120 KDa |
-Macromolecule #2: gp41
| Macromolecule | Name: gp41 / type: protein_or_peptide / ID: 2 / Name.synonym: SIV envelope transmembrane protein / Details: trimeric / Number of copies: 3 / Oligomeric state: trimer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Simian immunodeficiency virus / Strain: SIVMneCL8 / synonym: SIVmne / Cell: HuT 78 / Location in cell: viral membrane Simian immunodeficiency virus / Strain: SIVMneCL8 / synonym: SIVmne / Cell: HuT 78 / Location in cell: viral membrane |
| Molecular weight | Experimental: 41 KDa / Theoretical: 41 KDa |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.2 / Details: PBS |
| Staining | Type: NEGATIVE Details: Samples were mixed with BSA adsorbed 10 nm colloidal gold and vitrified for cryoelectron microscopy. |
| Grid | Details: holey carbon film |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: home built / Method: blot for two seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/ST |
|---|---|
| Temperature | Average: 77 K |
| Specialist optics | Energy filter - Name: GATAN GIF 2002 / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Details | We collected a tilt series |
| Date | Oct 1, 2005 |
| Image recording | Category: CCD / Film or detector model: GENERIC GATAN / Digitization - Sampling interval: 30 µm / Number real images: 3 / Average electron dose: 57 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 0.6 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 27500 |
| Sample stage | Specimen holder: eucentric / Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: 69.0 ° / Tilt series - Axis1 - Max angle: 63 ° |
- Image processing
Image processing
| Details | Average number of projections used in the 3D reconstructions: 2986. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 28.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: TOM Details: envelope complexes were averaged to give a final map of 28A resolution as determined by FSC at 0.5 cut-off. |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name: URO |
| Details | PDBEntryID_givenInChain. Protocol: rigid body. The gp120 monomer was manually fitted and three-fold symmetry was imposed during fitting with URO |
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT / Target criteria: quadratic misfit |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)