+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Organisation of a helical Bunyamwera virus ribonucleoprotein | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationhelical viral capsid / symbiont-mediated suppression of host gene expression / viral nucleocapsid / ribonucleoprotein complex / RNA binding Similarity search - Function | |||||||||
| Biological species |  Bunyamwera virus / Bunyamwera virus /  BUNV (virus) BUNV (virus) | |||||||||
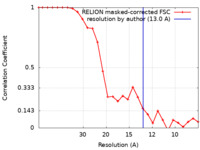
| Method | single particle reconstruction / cryo EM / Resolution: 13.0 Å | |||||||||
 Authors Authors | Hopkins FR / Fontana J / Barr JN | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: mBio / Year: 2022 Journal: mBio / Year: 2022Title: The Native Orthobunyavirus Ribonucleoprotein Possesses a Helical Architecture. Authors: Francis R Hopkins / Beatriz Álvarez-Rodríguez / George R Heath / Kyriakoulla Panayi / Samantha Hover / Thomas A Edwards / John N Barr / Juan Fontana /  Abstract: The order is the largest group of negative-sense RNA viruses, containing many lethal human pathogens for which approved anti-infective measures are not available. The bunyavirus genome consists of ...The order is the largest group of negative-sense RNA viruses, containing many lethal human pathogens for which approved anti-infective measures are not available. The bunyavirus genome consists of multiple negative-sense RNA segments enwrapped by the virus-encoded nucleocapsid protein (NP), which together with the viral polymerase form ribonucleoproteins (RNPs). RNPs represent substrates for RNA synthesis and virion assembly, which require inherent flexibility, consistent with the appearance of RNPs spilled from virions. These observations have resulted in conflicting models describing the overall RNP architecture. Here, we purified RNPs from Bunyamwera virus (BUNV), the prototypical orthobunyavirus. The lengths of purified RNPs imaged by negative staining resulted in 3 populations of RNPs, suggesting that RNPs possess a consistent method of condensation. Employing microscopy approaches, we conclusively show that the NP portion of BUNV RNPs is helical. Furthermore, we present a pseudo-atomic model for this portion based on a cryo-electron microscopy average at 13 Å resolution, which allowed us to fit the BUNV NP crystal structure by molecular dynamics. This model was confirmed by NP mutagenesis using a mini-genome system. The model shows that adjacent NP monomers in the RNP chain interact laterally through flexible N- and C-terminal arms only, with no longitudinal helix-stabilizing interactions, thus providing a potential model for the molecular basis for RNP flexibility. Excessive RNase treatment disrupts native RNPs, suggesting that RNA was key in maintaining the RNP structure. Overall, this work will inform studies on bunyaviral RNP assembly, packaging, and RNA replication, and aid in future antiviral strategies. Bunyaviruses are emerging RNA viruses that cause significant disease and economic burden and for which vaccines or therapies approved for humans are not available. The bunyavirus genome is wrapped up by the nucleoprotein (NP) and interacts with the viral polymerase, forming a ribonucleoprotein (RNP). This is the only form of the genome active for viral replication and assembly. However, until now how NPs are organized within an RNP was not known for any orthobunyavirus. Here, we purified RNPs from the prototypical orthobunyavirus, Bunyamwera virus, and employed microscopy approaches to show that the NP portion of the RNP was helical. We then combined our helical average with the known structure of an NP monomer, generating a pseudo-atomic model of this region. This arrangement allowed the RNPs to be highly flexible, which was critical for several stages of the viral replication cycle, such as segment circularization. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11847.map.gz emd_11847.map.gz | 735.8 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11847-v30.xml emd-11847-v30.xml emd-11847.xml emd-11847.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11847_fsc.xml emd_11847_fsc.xml | 2.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_11847.png emd_11847.png | 24.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11847 http://ftp.pdbj.org/pub/emdb/structures/EMD-11847 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11847 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11847 | HTTPS FTP |
-Validation report
| Summary document |  emd_11847_validation.pdf.gz emd_11847_validation.pdf.gz | 425.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11847_full_validation.pdf.gz emd_11847_full_validation.pdf.gz | 424.8 KB | Display | |
| Data in XML |  emd_11847_validation.xml.gz emd_11847_validation.xml.gz | 6.5 KB | Display | |
| Data in CIF |  emd_11847_validation.cif.gz emd_11847_validation.cif.gz | 7.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11847 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11847 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11847 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11847 | HTTPS FTP |
-Related structure data
| Related structure data |  7aoyMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11847.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11847.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 4.28 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Native ribonucleoprotein extracted from Bunyamwera virus
| Entire | Name: Native ribonucleoprotein extracted from Bunyamwera virus |
|---|---|
| Components |
|
-Supramolecule #1: Native ribonucleoprotein extracted from Bunyamwera virus
| Supramolecule | Name: Native ribonucleoprotein extracted from Bunyamwera virus type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Bunyamwera virus Bunyamwera virus |
-Macromolecule #1: Nucleoprotein
| Macromolecule | Name: Nucleoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 9 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  BUNV (virus) BUNV (virus) |
| Molecular weight | Theoretical: 26.69876 KDa |
| Sequence | String: MIELEFHDVA ANTSSTFDPE VAYANFKRVH TTGLSYDHIR IFYIKGREIK TSLAKRSEWE VTLNLGGWKI TVYNTNFPGN RNNPVPDDG LTLHRLSGFL ARYLLEKMLK VSEPEKLIIK SKIINPLAEK NGITWNDGEE VYLSFFPGSE MFLGTFRFYP L AIGIYKVQ ...String: MIELEFHDVA ANTSSTFDPE VAYANFKRVH TTGLSYDHIR IFYIKGREIK TSLAKRSEWE VTLNLGGWKI TVYNTNFPGN RNNPVPDDG LTLHRLSGFL ARYLLEKMLK VSEPEKLIIK SKIINPLAEK NGITWNDGEE VYLSFFPGSE MFLGTFRFYP L AIGIYKVQ RKEMEPKYLE KTMRQRYMGL EAATWTVSKL TEVQSALTVV SSLGWKKTNV SAAARDFLAK FGINM |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: NITROGEN / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 9.0 sec. / Average electron dose: 51.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 130000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller