+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10444 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Sub-tomogram average of the Sulfolobus acidocaldarius S-layer | |||||||||
 Map data Map data | Sub-tomogram average of the Sulfolobus acidocaldarius S-layer | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Sulfolobus acidocaldarius (acidophilic) Sulfolobus acidocaldarius (acidophilic) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 23.0 Å | |||||||||
 Authors Authors | Daum B / Gambelli L | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: Architecture and modular assembly of S-layers revealed by electron cryotomography. Authors: Lavinia Gambelli / Benjamin H Meyer / Mathew McLaren / Kelly Sanders / Tessa E F Quax / Vicki A M Gold / Sonja-Verena Albers / Bertram Daum /   Abstract: Surface protein layers (S-layers) often form the only structural component of the archaeal cell wall and are therefore important for cell survival. S-layers have a plethora of cellular functions ...Surface protein layers (S-layers) often form the only structural component of the archaeal cell wall and are therefore important for cell survival. S-layers have a plethora of cellular functions including maintenance of cell shape, osmotic, and mechanical stability, the formation of a semipermeable protective barrier around the cell, and cell-cell interaction, as well as surface adhesion. Despite the central importance of S-layers for archaeal life, their 3-dimensional (3D) architecture is still poorly understood. Here we present detailed 3D electron cryomicroscopy maps of archaeal S-layers from 3 different strains. We were able to pinpoint the positions and determine the structure of the 2 subunits SlaA and SlaB. We also present a model describing the assembly of the mature S-layer. #1:  Journal: To Be Published Journal: To Be PublishedTitle: Architecture and modular assembly of Sulfolobus S-layers revealed by electron cryo-tomography Authors: Gambelli L / Meyer BH / McLaren M / Sanders K / Quax TEF / Gold VAM / Albers SVA / Daum B | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10444.map.gz emd_10444.map.gz | 5.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10444-v30.xml emd-10444-v30.xml emd-10444.xml emd-10444.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10444.png emd_10444.png | 55.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10444 http://ftp.pdbj.org/pub/emdb/structures/EMD-10444 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10444 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10444 | HTTPS FTP |
-Validation report
| Summary document |  emd_10444_validation.pdf.gz emd_10444_validation.pdf.gz | 245.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10444_full_validation.pdf.gz emd_10444_full_validation.pdf.gz | 244.4 KB | Display | |
| Data in XML |  emd_10444_validation.xml.gz emd_10444_validation.xml.gz | 4.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10444 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10444 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10444 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10444 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10444.map.gz / Format: CCP4 / Size: 5.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10444.map.gz / Format: CCP4 / Size: 5.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sub-tomogram average of the Sulfolobus acidocaldarius S-layer | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.70655 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
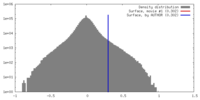
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SlaA/SlaB array
| Entire | Name: SlaA/SlaB array |
|---|---|
| Components |
|
-Supramolecule #1: SlaA/SlaB array
| Supramolecule | Name: SlaA/SlaB array / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Sulfolobus acidocaldarius (acidophilic) Sulfolobus acidocaldarius (acidophilic) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Buffer | pH: 3 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Specialist optics | Energy filter - Slit width: 40 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.33 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Cooling holder cryogen: NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 23.0 Å / Resolution method: DIFFRACTION PATTERN/LAYERLINES / Software - Name: PEET / Number subtomograms used: 325 |
|---|---|
| Extraction | Number tomograms: 1 / Number images used: 325 |
| CTF correction | Software - Name:  IMOD IMOD |
| Final angle assignment | Type: OTHER / Software - Name: PEET |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)