[English] 日本語
 Yorodumi
Yorodumi- EMDB-8153: Masked monomer of ATP synthase dimer from Yarrowia lipolytica, su... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8153 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Masked monomer of ATP synthase dimer from Yarrowia lipolytica, subclass 1 | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Yarrowia lipolytica (yeast) Yarrowia lipolytica (yeast) | |||||||||
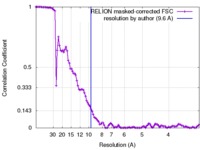
| Method | single particle reconstruction / cryo EM / Resolution: 9.6 Å | |||||||||
 Authors Authors | Hahn A / Parey K / Bublitz M / Mills DJ / Zickermann V / Vonck J / Kuehlbrandt W / Meier T | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2016 Journal: Mol Cell / Year: 2016Title: Structure of a Complete ATP Synthase Dimer Reveals the Molecular Basis of Inner Mitochondrial Membrane Morphology. Authors: Alexander Hahn / Kristian Parey / Maike Bublitz / Deryck J Mills / Volker Zickermann / Janet Vonck / Werner Kühlbrandt / Thomas Meier /  Abstract: We determined the structure of a complete, dimeric F1Fo-ATP synthase from yeast Yarrowia lipolytica mitochondria by a combination of cryo-EM and X-ray crystallography. The final structure resolves 58 ...We determined the structure of a complete, dimeric F1Fo-ATP synthase from yeast Yarrowia lipolytica mitochondria by a combination of cryo-EM and X-ray crystallography. The final structure resolves 58 of the 60 dimer subunits. Horizontal helices of subunit a in Fo wrap around the c-ring rotor, and a total of six vertical helices assigned to subunits a, b, f, i, and 8 span the membrane. Subunit 8 (A6L in human) is an evolutionary derivative of the bacterial b subunit. On the lumenal membrane surface, subunit f establishes direct contact between the two monomers. Comparison with a cryo-EM map of the F1Fo monomer identifies subunits e and g at the lateral dimer interface. They do not form dimer contacts but enable dimer formation by inducing a strong membrane curvature of ∼100°. Our structure explains the structural basis of cristae formation in mitochondria, a landmark signature of eukaryotic cell morphology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8153.map.gz emd_8153.map.gz | 93.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8153-v30.xml emd-8153-v30.xml emd-8153.xml emd-8153.xml | 11.9 KB 11.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8153_fsc.xml emd_8153_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_8153.png emd_8153.png | 29.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8153 http://ftp.pdbj.org/pub/emdb/structures/EMD-8153 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8153 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8153 | HTTPS FTP |
-Validation report
| Summary document |  emd_8153_validation.pdf.gz emd_8153_validation.pdf.gz | 79.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8153_full_validation.pdf.gz emd_8153_full_validation.pdf.gz | 78.4 KB | Display | |
| Data in XML |  emd_8153_validation.xml.gz emd_8153_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8153 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8153 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8153 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8153 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8153.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8153.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.63 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Fo ATP synthase dimer
| Entire | Name: Fo ATP synthase dimer |
|---|---|
| Components |
|
-Supramolecule #1: Fo ATP synthase dimer
| Supramolecule | Name: Fo ATP synthase dimer / type: complex / ID: 1 / Parent: 0 Details: The Fo section of the F1Fo dimer was masked for refinement |
|---|---|
| Source (natural) | Organism:  Yarrowia lipolytica (yeast) Yarrowia lipolytica (yeast) |
| Molecular weight | Theoretical: 1.2 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 30 mM MOPS-NaOH pH 7.5, 2 mM MgCl2, 0.5 mM EDTA, 50 mM NaCl, 0.05% (w/v) digitonin |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV / Details: blotting for 7 to 9 s. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Sampling interval: 5.0 µm / Digitization - Frames/image: 2-21 / Number real images: 2500 / Average exposure time: 6.0 sec. / Average electron dose: 18.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 30675 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 4.2 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 20000 |
| Sample stage | Specimen holder model: JEOL 3200FSC CRYOHOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller