+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8r1a | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
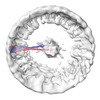
| Title | Model of the membrane-bound GBP1 oligomer | |||||||||
 Components Components | Guanylate binding protein 1 | |||||||||
 Keywords Keywords |  ANTIMICROBIAL PROTEIN / ANTIMICROBIAL PROTEIN /  Oligomer / Oligomer /  GTPase / Interferon-induced GTPase / Interferon-induced | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  ELECTRON MICROSCOPY / subtomogram averaging / ELECTRON MICROSCOPY / subtomogram averaging /  cryo EM / Resolution: 26.8 Å cryo EM / Resolution: 26.8 Å | |||||||||
 Authors Authors | Weismehl, M. / Chu, X. / Kutsch, M. / Lauterjung, P. / Herrmann, C. / Kudryashev, M. / Daumke, O. | |||||||||
| Funding support |  Germany, 2items Germany, 2items
| |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2024 Journal: EMBO J / Year: 2024Title: Structural insights into the activation mechanism of antimicrobial GBP1. Authors: Marius Weismehl / Xiaofeng Chu / Miriam Kutsch / Paul Lauterjung / Christian Herrmann / Misha Kudryashev / Oliver Daumke /   Abstract: The dynamin-related human guanylate-binding protein 1 (GBP1) mediates host defenses against microbial pathogens. Upon GTP binding and hydrolysis, auto-inhibited GBP1 monomers dimerize and assemble ...The dynamin-related human guanylate-binding protein 1 (GBP1) mediates host defenses against microbial pathogens. Upon GTP binding and hydrolysis, auto-inhibited GBP1 monomers dimerize and assemble into soluble and membrane-bound oligomers, which are crucial for innate immune responses. How higher-order GBP1 oligomers are built from dimers, and how assembly is coordinated with nucleotide-dependent conformational changes, has remained elusive. Here, we present cryo-electron microscopy-based structural data of soluble and membrane-bound GBP1 oligomers, which show that GBP1 assembles in an outstretched dimeric conformation. We identify a surface-exposed helix in the large GTPase domain that contributes to the oligomerization interface, and we probe its nucleotide- and dimerization-dependent movements that facilitate the formation of an antimicrobial protein coat on a gram-negative bacterial pathogen. Our results reveal a sophisticated activation mechanism for GBP1, in which nucleotide-dependent structural changes coordinate dimerization, oligomerization, and membrane binding to allow encapsulation of pathogens within an antimicrobial protein coat. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8r1a.cif.gz 8r1a.cif.gz | 601.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8r1a.ent.gz pdb8r1a.ent.gz | 485.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8r1a.json.gz 8r1a.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/r1/8r1a https://data.pdbj.org/pub/pdb/validation_reports/r1/8r1a ftp://data.pdbj.org/pub/pdb/validation_reports/r1/8r1a ftp://data.pdbj.org/pub/pdb/validation_reports/r1/8r1a | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  18806MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 69267.930 Da / Num. of mol.: 6 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) / References: UniProt: Q5D1D5 Escherichia coli (E. coli) / References: UniProt: Q5D1D5#2: Chemical | ChemComp-AF3 /  Aluminium fluoride Aluminium fluoride#3: Chemical | ChemComp-MG / #4: Chemical | ChemComp-GDP /  Guanosine diphosphate Guanosine diphosphateHas ligand of interest | N | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: subtomogram averaging |
- Sample preparation
Sample preparation
| Component | Name: Membrane-bound oligomer of human guanylate-binding protein 1 (GBP1) Type: COMPLEX / Details: in complex with GDP-AlFx / Entity ID: #1 / Source: RECOMBINANT |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Buffer solution | pH: 7.9 Details: 50 mM Tris-HCl, 150 mM NaCl, 5 mM MgCl2 (supplemented with 200 uM GDP, 300 uM AlCl3, 10 mM NaF) |
| Specimen | Conc.: 0.68 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal magnification: 42000 X / Nominal defocus max: 5000 nm / Nominal defocus min: 2000 nm Bright-field microscopy / Nominal magnification: 42000 X / Nominal defocus max: 5000 nm / Nominal defocus min: 2000 nm |
| Image recording | Electron dose: 2.8 e/Å2 / Avg electron dose per subtomogram: 130 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) Details: Tilt series were acquired with a Hybrid-STA (Sanchez et al. 2020, Nat Commun): exposure dose of 2.8 e-/A2 for non-zero tilted projection and 14.4 e-/A2 for zero tilted projection |
- Processing
Processing
CTF correction | Type: PHASE FLIPPING ONLY |
|---|---|
| Symmetry | Point symmetry : C1 (asymmetric) : C1 (asymmetric) |
3D reconstruction | Resolution: 26.8 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 6146 / Symmetry type: POINT |
| EM volume selection | Num. of tomograms: 104 / Num. of volumes extracted: 70160 |
 Movie
Movie Controller
Controller



 PDBj
PDBj







