[English] 日本語
 Yorodumi
Yorodumi- PDB-7vq0: Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) boun... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7vq0 | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
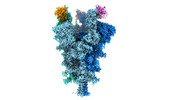
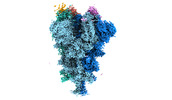
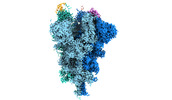
| Title | Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing nanobodies P86 | |||||||||||||||||||||||||||
 Components Components |
| |||||||||||||||||||||||||||
 Keywords Keywords |  VIRAL PROTEIN/IMMUNE SYSTEM / spike / VIRAL PROTEIN/IMMUNE SYSTEM / spike /  nanobody / VHH / nanobody / VHH /  VIRAL PROTEIN / VIRAL PROTEIN /  VIRAL PROTEIN-IMMUNE SYSTEM complex VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry /  membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell /  receptor ligand activity / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / receptor ligand activity / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane /  viral envelope / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / viral envelope / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||||||||||||||||||||
| Biological species |   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2  Vicugna pacos (alpaca) Vicugna pacos (alpaca) | |||||||||||||||||||||||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.03 Å cryo EM / Resolution: 3.03 Å | |||||||||||||||||||||||||||
 Authors Authors | Maeda, R. / Fujita, J. / Konishi, Y. / Kazuma, Y. / Yamazaki, H. / Anzai, I. / Yamaguchi, K. / Kasai, K. / Nagata, K. / Yamaoka, Y. ...Maeda, R. / Fujita, J. / Konishi, Y. / Kazuma, Y. / Yamazaki, H. / Anzai, I. / Yamaguchi, K. / Kasai, K. / Nagata, K. / Yamaoka, Y. / Miyakawa, K. / Ryo, A. / Shirakawa, K. / Makino, F. / Matsuura, Y. / Inoue, T. / Imura, A. / Namba, K. / Takaori-Kondo, A. | |||||||||||||||||||||||||||
| Funding support |  Japan, 8items Japan, 8items
| |||||||||||||||||||||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2022 Journal: Commun Biol / Year: 2022Title: A panel of nanobodies recognizing conserved hidden clefts of all SARS-CoV-2 spike variants including Omicron. Authors: Ryota Maeda / Junso Fujita / Yoshinobu Konishi / Yasuhiro Kazuma / Hiroyuki Yamazaki / Itsuki Anzai / Tokiko Watanabe / Keishi Yamaguchi / Kazuki Kasai / Kayoko Nagata / Yutaro Yamaoka / Kei ...Authors: Ryota Maeda / Junso Fujita / Yoshinobu Konishi / Yasuhiro Kazuma / Hiroyuki Yamazaki / Itsuki Anzai / Tokiko Watanabe / Keishi Yamaguchi / Kazuki Kasai / Kayoko Nagata / Yutaro Yamaoka / Kei Miyakawa / Akihide Ryo / Kotaro Shirakawa / Kei Sato / Fumiaki Makino / Yoshiharu Matsuura / Tsuyoshi Inoue / Akihiro Imura / Keiichi Namba / Akifumi Takaori-Kondo /  Abstract: We are amid the historic coronavirus infectious disease 2019 (COVID-19) pandemic. Imbalances in the accessibility of vaccines, medicines, and diagnostics among countries, regions, and populations, ...We are amid the historic coronavirus infectious disease 2019 (COVID-19) pandemic. Imbalances in the accessibility of vaccines, medicines, and diagnostics among countries, regions, and populations, and those in war crises, have been problematic. Nanobodies are small, stable, customizable, and inexpensive to produce. Herein, we present a panel of nanobodies that can detect the spike proteins of five SARS-CoV-2 variants of concern (VOCs) including Omicron. Here we show via ELISA, lateral flow, kinetic, flow cytometric, microscopy, and Western blotting assays that our nanobodies can quantify the spike variants. This panel of nanobodies broadly neutralizes viral infection caused by pseudotyped and authentic SARS-CoV-2 VOCs. Structural analyses show that the P86 clone targets epitopes that are conserved yet unclassified on the receptor-binding domain (RBD) and contacts the N-terminal domain (NTD). Human antibodies rarely access both regions; consequently, the clone buries hidden crevasses of SARS-CoV-2 spike proteins that go undetected by conventional antibodies. | |||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7vq0.cif.gz 7vq0.cif.gz | 650.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7vq0.ent.gz pdb7vq0.ent.gz | 542.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7vq0.json.gz 7vq0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/vq/7vq0 https://data.pdbj.org/pub/pdb/validation_reports/vq/7vq0 ftp://data.pdbj.org/pub/pdb/validation_reports/vq/7vq0 ftp://data.pdbj.org/pub/pdb/validation_reports/vq/7vq0 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  32078MC  7vpyC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein / Antibody , 2 types, 6 molecules ABCDEF
| #1: Protein |  Spike protein / S glycoprotein / E2 / Peplomer protein Spike protein / S glycoprotein / E2 / Peplomer proteinMass: 138154.984 Da / Num. of mol.: 3 / Mutation: D614G, R682G, R683S, R685S, K986P, V987P Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2Gene: S, 2 / Production host:   Homo sapiens (human) / References: UniProt: P0DTC2 Homo sapiens (human) / References: UniProt: P0DTC2#2: Antibody | Mass: 13432.936 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Vicugna pacos (alpaca) / Production host: Vicugna pacos (alpaca) / Production host:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
|---|
-Sugars , 4 types, 56 molecules 
| #3: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose  / Mass: 424.401 Da / Num. of mol.: 17 / Mass: 424.401 Da / Num. of mol.: 17Source method: isolated from a genetically manipulated source #4: Polysaccharide |  / Mass: 570.542 Da / Num. of mol.: 3 / Mass: 570.542 Da / Num. of mol.: 3Source method: isolated from a genetically manipulated source #5: Polysaccharide | alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta- ...alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose  / Mass: 586.542 Da / Num. of mol.: 6 / Mass: 586.542 Da / Num. of mol.: 6Source method: isolated from a genetically manipulated source #6: Sugar | ChemComp-NAG /  N-Acetylglucosamine N-Acetylglucosamine |
|---|
-Details
| Has ligand of interest | N |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular weight | Experimental value: NO | ||||||||||||||||||||||||
| Source (natural) |
| ||||||||||||||||||||||||
| Source (recombinant) |
| ||||||||||||||||||||||||
| Buffer solution | pH: 7.4 | ||||||||||||||||||||||||
| Buffer component |
| ||||||||||||||||||||||||
| Specimen | Conc.: 0.1 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES / Details: Mixed with 5 times molar excess of P86. : YES / Details: Mixed with 5 times molar excess of P86. | ||||||||||||||||||||||||
| Specimen support | Details: The graphene grid was chemically oxidized and modified. Grid material: GOLD / Grid mesh size: 200 divisions/in. / Grid type: Quantifoil R1.2/1.3 | ||||||||||||||||||||||||
Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: JEOL CRYO ARM 300 |
|---|---|
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal magnification: 60000 X / Nominal defocus max: 2000 nm / Nominal defocus min: 500 nm / Cs Bright-field microscopy / Nominal magnification: 60000 X / Nominal defocus max: 2000 nm / Nominal defocus min: 500 nm / Cs : 2.7 mm : 2.7 mm |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: JEOL CRYOSPECPORTER |
| Image recording | Average exposure time: 3 sec. / Electron dose: 60 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) / Num. of grids imaged: 1 |
| EM imaging optics | Energyfilter name : In-column Omega Filter / Energyfilter slit width: 20 eV : In-column Omega Filter / Energyfilter slit width: 20 eV |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.19.1_4122: / Classification: refinement | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||||||||||||||
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 878975 | ||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry : C1 (asymmetric) : C1 (asymmetric) | ||||||||||||||||||||||||||||||||||||
3D reconstruction | Resolution: 3.03 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 38004 / Algorithm: FOURIER SPACE / Symmetry type: POINT | ||||||||||||||||||||||||||||||||||||
| Atomic model building | PDB-ID: 7KRR | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller


 PDBj
PDBj







