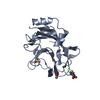
Entry Database : PDB / ID : 5wxfTitle Crystal structure of uPA in complex with upain-2-2 Urokinase-type plasminogen activator chain B upain-2-2 peptide Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Phage display vector pTDisp (others) Method / / / Resolution : 1.46 Å Authors Jiang, L. / Huang, M. Funding support Organization Grant number Country NSFC 31400637
Journal : Biochim. Biophys. Acta / Year : 2018Title : Cleavage of peptidic inhibitors by target protease is caused by peptide conformational transition.Authors : Jiang, L. / Oldenburg, E. / Kromann-Hansen, T. / Xu, P. / Jensen, J.K. / Andreasen, P.A. / Huang, M. History Deposition Jan 7, 2017 Deposition site / Processing site Revision 1.0 Jul 11, 2018 Provider / Type Revision 1.1 Jul 18, 2018 Group / Database references / Category / citation_authorItem _citation.country / _citation.journal_abbrev ... _citation.country / _citation.journal_abbrev / _citation.journal_id_ASTM / _citation.journal_id_CSD / _citation.journal_id_ISSN / _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.pdbx_database_id_DOI / _citation.pdbx_database_id_PubMed / _citation.title / _citation.year Revision 1.2 Nov 22, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_conn Item / _database_2.pdbx_database_accession / _struct_conn.pdbx_leaving_atom_flag
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Serine proteases / uPA / pepetide inhibitors / HYDROLASE-HYDROLASE INHIBITOR complex
Serine proteases / uPA / pepetide inhibitors / HYDROLASE-HYDROLASE INHIBITOR complex Function and homology information
Function and homology information u-plasminogen activator / regulation of smooth muscle cell-matrix adhesion / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation /
u-plasminogen activator / regulation of smooth muscle cell-matrix adhesion / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation /  regulation of fibrinolysis /
regulation of fibrinolysis /  regulation of wound healing / protein complex involved in cell-matrix adhesion / negative regulation of plasminogen activation / regulation of smooth muscle cell migration /
regulation of wound healing / protein complex involved in cell-matrix adhesion / negative regulation of plasminogen activation / regulation of smooth muscle cell migration /  regulation of signaling receptor activity ...
regulation of signaling receptor activity ... u-plasminogen activator / regulation of smooth muscle cell-matrix adhesion / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation /
u-plasminogen activator / regulation of smooth muscle cell-matrix adhesion / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation /  regulation of fibrinolysis /
regulation of fibrinolysis /  regulation of wound healing / protein complex involved in cell-matrix adhesion / negative regulation of plasminogen activation / regulation of smooth muscle cell migration /
regulation of wound healing / protein complex involved in cell-matrix adhesion / negative regulation of plasminogen activation / regulation of smooth muscle cell migration /  regulation of signaling receptor activity / serine-type endopeptidase complex / Dissolution of Fibrin Clot / smooth muscle cell migration / plasminogen activation / regulation of cell adhesion mediated by integrin / tertiary granule membrane / negative regulation of fibrinolysis /
regulation of signaling receptor activity / serine-type endopeptidase complex / Dissolution of Fibrin Clot / smooth muscle cell migration / plasminogen activation / regulation of cell adhesion mediated by integrin / tertiary granule membrane / negative regulation of fibrinolysis /  regulation of cell adhesion / specific granule membrane /
regulation of cell adhesion / specific granule membrane /  serine protease inhibitor complex /
serine protease inhibitor complex /  fibrinolysis /
fibrinolysis /  chemotaxis /
chemotaxis /  blood coagulation / regulation of cell population proliferation / response to hypoxia / positive regulation of cell migration / external side of plasma membrane / serine-type endopeptidase activity /
blood coagulation / regulation of cell population proliferation / response to hypoxia / positive regulation of cell migration / external side of plasma membrane / serine-type endopeptidase activity /  focal adhesion / Neutrophil degranulation /
focal adhesion / Neutrophil degranulation /  cell surface /
cell surface /  signal transduction /
signal transduction /  proteolysis /
proteolysis /  extracellular space / extracellular exosome / extracellular region /
extracellular space / extracellular exosome / extracellular region /  plasma membrane
plasma membrane
 Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.46 Å
MOLECULAR REPLACEMENT / Resolution: 1.46 Å  Authors
Authors China, 1items
China, 1items  Citation
Citation Journal: Biochim. Biophys. Acta / Year: 2018
Journal: Biochim. Biophys. Acta / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5wxf.cif.gz
5wxf.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5wxf.ent.gz
pdb5wxf.ent.gz PDB format
PDB format 5wxf.json.gz
5wxf.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/wx/5wxf
https://data.pdbj.org/pub/pdb/validation_reports/wx/5wxf ftp://data.pdbj.org/pub/pdb/validation_reports/wx/5wxf
ftp://data.pdbj.org/pub/pdb/validation_reports/wx/5wxf


 Links
Links Assembly
Assembly
 Components
Components
 Homo sapiens (human) / Gene: PLAU / Production host:
Homo sapiens (human) / Gene: PLAU / Production host: 
 Komagataella pastoris (fungus) / References: UniProt: P00749,
Komagataella pastoris (fungus) / References: UniProt: P00749,  u-plasminogen activator
u-plasminogen activator Sulfate
Sulfate Water
Water X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRF
SSRF  / Beamline: BL17U / Wavelength: 0.979 Å
/ Beamline: BL17U / Wavelength: 0.979 Å : 0.979 Å / Relative weight: 1
: 0.979 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller
















 PDBj
PDBj



