[English] 日本語
 Yorodumi
Yorodumi- PDB-3qz0: Structure of Treponema denticola Factor H Binding protein (FhbB),... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3qz0 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of Treponema denticola Factor H Binding protein (FhbB), selenomethionine derivative | ||||||
 Components Components | Factor H binding protein | ||||||
 Keywords Keywords |  PROTEIN BINDING / Factor H binding protein / PROTEIN BINDING / Factor H binding protein /  Factor H Factor H | ||||||
| Function / homology |  Function and homology information Function and homology informationSingle alpha-helices involved in coiled-coils or other helix-helix interfaces - #2300 / : / : / Factor H binding protein B / Single alpha-helices involved in coiled-coils or other helix-helix interfaces / Helix non-globular / Special / Prokaryotic membrane lipoprotein lipid attachment site profile. Similarity search - Domain/homology | ||||||
| Biological species |   Treponema denticola (bacteria) Treponema denticola (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.77 Å SAD / Resolution: 1.77 Å | ||||||
 Authors Authors | Miller, D.P. / McDowell, J.V. / Heroux, A. / Bell, J.K. / Marconi, R.T. / Conrad, D.H. / Burgner, J.W. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2012 Journal: J.Biol.Chem. / Year: 2012Title: Structure of factor H-binding protein B (FhbB) of the periopathogen, Treponema denticola: insights into progression of periodontal disease. Authors: Miller, D.P. / Bell, J.K. / McDowell, J.V. / Conrad, D.H. / Burgner, J.W. / Heroux, A. / Marconi, R.T. #1: Journal: Acta Crystallogr.,Sect.F / Year: 2011 Title: Crystallization of the factor H-binding protein, FhbB, from the periopathogen Treponema denticola. Authors: Miller, D.P. / McDowell, J.V. / Bell, J.K. / Marconi, R.T. #2:  Journal: Infect.Immun. / Year: 2009 Journal: Infect.Immun. / Year: 2009Title: Analysis of unique interaction between complement regulatory protein factor H and periodontal pathogen Treponema denticola Authors: McDowell, J.V. / Huang, B. / Fenno, J.C. / Marconi, R.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3qz0.cif.gz 3qz0.cif.gz | 43.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3qz0.ent.gz pdb3qz0.ent.gz | 34.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3qz0.json.gz 3qz0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qz/3qz0 https://data.pdbj.org/pub/pdb/validation_reports/qz/3qz0 ftp://data.pdbj.org/pub/pdb/validation_reports/qz/3qz0 ftp://data.pdbj.org/pub/pdb/validation_reports/qz/3qz0 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 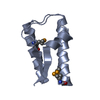
| ||||||||||||
| 2 | 
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 10700.863 Da / Num. of mol.: 2 / Fragment: residues 24-102 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Treponema denticola (bacteria) / Strain: 35405 / Gene: fhbB, TDE_0108 / Plasmid: pET46 Ek/LIC / Production host: Treponema denticola (bacteria) / Strain: 35405 / Gene: fhbB, TDE_0108 / Plasmid: pET46 Ek/LIC / Production host:   Escherichia coli (E. coli) / Strain (production host): B834(DE3)(pLyS) / References: UniProt: Q73RI0 Escherichia coli (E. coli) / Strain (production host): B834(DE3)(pLyS) / References: UniProt: Q73RI0#2: Chemical |  Thiocyanate Thiocyanate#3: Chemical | ChemComp-GOL / |  Glycerol Glycerol#4: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.2 Å3/Da / Density % sol: 44.08 % |
|---|---|
Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 1.39 M sodium citrate, 0.1 M HEPES, 0.2 M sodium thiocyanate, 10 mM dithiothreitol, 10% glycerol, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X25 / Wavelength: 1.2822 Å / Beamline: X25 / Wavelength: 1.2822 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: May 28, 2010 |
| Radiation | Monochromator: Double silicon(111) crystal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.2822 Å / Relative weight: 1 : 1.2822 Å / Relative weight: 1 |
| Reflection | Resolution: 1.77→50 Å / Num. all: 19293 / Num. obs: 19167 / % possible obs: 99.3 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 1 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  SAD / Resolution: 1.77→45.207 Å / SU ML: 0.18 / σ(F): 0.24 / Stereochemistry target values: MLHL SAD / Resolution: 1.77→45.207 Å / SU ML: 0.18 / σ(F): 0.24 / Stereochemistry target values: MLHL
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 41.98 Å2 / ksol: 0.387 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.77→45.207 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller










 PDBj
PDBj










