+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3ov5 | ||||||
|---|---|---|---|---|---|---|---|
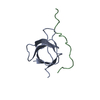
| Title | Atomic structure of the Xanthomonas citri VirB7 globular domain. | ||||||
 Components Components | Uncharacterized protein | ||||||
 Keywords Keywords |  PROTEIN TRANSPORT / PROTEIN TRANSPORT /  Type IV secretion system component / VirB7 (XAC2622) / Type IV secretion system component / VirB7 (XAC2622) /  Bacterial outer membrane / XANTHOMONAS AXONOPODIS PV CITRI Bacterial outer membrane / XANTHOMONAS AXONOPODIS PV CITRI | ||||||
| Function / homology | Phage tail protein beta-alpha-beta fold - #70 / Toxin co-regulated pilus biosynthesis protein Q, C-terminal / Toxin co-regulated pilus biosynthesis protein Q / Phage tail protein beta-alpha-beta fold / 3-Layer(bab) Sandwich / Alpha Beta /  ISOPROPYL ALCOHOL / Toxin co-regulated pilus biosynthesis protein Q C-terminal domain-containing protein ISOPROPYL ALCOHOL / Toxin co-regulated pilus biosynthesis protein Q C-terminal domain-containing protein Function and homology information Function and homology information | ||||||
| Biological species |   Xanthomonas axonopodis pv. citri (bacteria) Xanthomonas axonopodis pv. citri (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.04 Å MOLECULAR REPLACEMENT / Resolution: 1.04 Å | ||||||
 Authors Authors | Souza, D.P. / Farah, C.S. | ||||||
 Citation Citation |  Journal: Plos Pathog. / Year: 2011 Journal: Plos Pathog. / Year: 2011Title: A Component of the Xanthomonadaceae Type IV Secretion System Combines a VirB7 Motif with a N0 Domain Found in Outer Membrane Transport Proteins. Authors: Souza, D.P. / Andrade, M.O. / Alvarez-Martinez, C.E. / Arantes, G.M. / Farah, C.S. / Salinas, R.K. | ||||||
| History |
| ||||||
| Remark 999 | MET A 50 is INITIATING METHIONINE and CLONING ARTIFACT. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3ov5.cif.gz 3ov5.cif.gz | 50.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3ov5.ent.gz pdb3ov5.ent.gz | 36.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3ov5.json.gz 3ov5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ov/3ov5 https://data.pdbj.org/pub/pdb/validation_reports/ov/3ov5 ftp://data.pdbj.org/pub/pdb/validation_reports/ov/3ov5 ftp://data.pdbj.org/pub/pdb/validation_reports/ov/3ov5 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 9021.133 Da / Num. of mol.: 1 / Fragment: Globular Domain, residues 51-134 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Xanthomonas axonopodis pv. citri (bacteria) Xanthomonas axonopodis pv. citri (bacteria)Strain: 306 / Gene: XAC2622 / Plasmid: pET-11a / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3)RP / References: UniProt: Q8PJB3 Escherichia coli (E. coli) / Strain (production host): BL21(DE3)RP / References: UniProt: Q8PJB3 |
|---|---|
| #2: Chemical | ChemComp-IPA /  Isopropyl alcohol Isopropyl alcohol |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.78 Å3/Da / Density % sol: 31.05 % |
|---|---|
Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 7.5 Details: 14 mg/mL protein in 5 mM Tris-Cl pH 7.5 and 25 mM sodium chloride was submitted to vapor diffusion sitting-drop crystallization trials at 291 K. Large plates appeared after one day over a ...Details: 14 mg/mL protein in 5 mM Tris-Cl pH 7.5 and 25 mM sodium chloride was submitted to vapor diffusion sitting-drop crystallization trials at 291 K. Large plates appeared after one day over a reservoir solution comprising 1.4 M ammonium sulphate and 4 % (v/v) isopropyl alcohol. Reservoir solution supplemented with 25 % (v/v) glycerol was used as cryoprotectant. , VAPOR DIFFUSION, SITTING DROP |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  LNLS LNLS  / Beamline: W01B-MX2 / Wavelength: 0.9537 Å / Beamline: W01B-MX2 / Wavelength: 0.9537 Å |
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Sep 30, 2009 |
| Radiation | Monochromator: Vertical collimator mirror + double Si(111) crystal monochromator + toroidal focus mirror Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9537 Å / Relative weight: 1 : 0.9537 Å / Relative weight: 1 |
| Reflection | Resolution: 1.04→30 Å / Num. all: 30386 / Num. obs: 30320 / % possible obs: 96.3 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 12.1 % / Biso Wilson estimate: 5.4 Å2 / Rmerge(I) obs: 0.068 / Net I/σ(I): 34.8 |
| Reflection shell | Resolution: 1.04→1.08 Å / Redundancy: 8.9 % / Rmerge(I) obs: 0.293 / Mean I/σ(I) obs: 5.1 / Num. unique all: 2449 / % possible all: 79 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: Globular domain from NMR structure of the same protein. Resolution: 1.04→23.81 Å / Cor.coef. Fo:Fc: 0.977 / Cor.coef. Fo:Fc free: 0.97 / SU B: 0.674 / SU ML: 0.016 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R Free: 0.027 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: BABINET MODEL WITH MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 10.366 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.04→23.81 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.04→1.067 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller













 PDBj
PDBj



