+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2xiw | ||||||
|---|---|---|---|---|---|---|---|
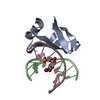
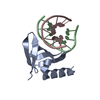
| Title | Crystal structure of the Sac7d-derived IgG1-binder C3-C24S | ||||||
 Components Components | DNA-BINDING PROTEIN 7D | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN DNA BINDING PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |    SULFOLOBUS ACIDOCALDARIUS (acidophilic) SULFOLOBUS ACIDOCALDARIUS (acidophilic) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.5 Å SAD / Resolution: 1.5 Å | ||||||
 Authors Authors | Bellinzoni, M. / Colinet, S. / Behar, G. / Alzari, P.M. / Pecorari, F. | ||||||
 Citation Citation |  Journal: Protein Eng.Des.Sel. / Year: 2013 Journal: Protein Eng.Des.Sel. / Year: 2013Title: Tolerance of the Archaeal Sac7D Scaffold Protein to Alternative Library Designs: Characterization of Anti-Immunoglobulin G Affitins. Authors: Behar, G. / Bellinzoni, M. / Maillasson, M. / Paillard-Laurance, L. / Alzari, P.M. / He, X. / Mouratou, B. / Pecorari, F. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2xiw.cif.gz 2xiw.cif.gz | 70.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2xiw.ent.gz pdb2xiw.ent.gz | 56.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2xiw.json.gz 2xiw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xi/2xiw https://data.pdbj.org/pub/pdb/validation_reports/xi/2xiw ftp://data.pdbj.org/pub/pdb/validation_reports/xi/2xiw ftp://data.pdbj.org/pub/pdb/validation_reports/xi/2xiw | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
| ||||||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.71044, 0.63121, 0.31122), Vector  : : |
- Components
Components
| #1: Protein | Mass: 9168.037 Da / Num. of mol.: 2 / Mutation: YES Source method: isolated from a genetically manipulated source Source: (gene. exp.)    SULFOLOBUS ACIDOCALDARIUS (acidophilic) SULFOLOBUS ACIDOCALDARIUS (acidophilic)Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): DH5[ALPHA] / Variant (production host): IQ / References: UniProt: P13123 ESCHERICHIA COLI (E. coli) / Strain (production host): DH5[ALPHA] / Variant (production host): IQ / References: UniProt: P13123#2: Chemical |  Sulfate Sulfate#3: Chemical | ChemComp-CL / |  Chloride Chloride#4: Water | ChemComp-HOH / |  Water WaterSequence details | N-TERMINAL STRETCH MRGSHHHHHH | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.92 Å3/Da / Density % sol: 57.9 % / Description: NONE |
|---|---|
Crystal grow | Details: 3.5 M (NH4)2SO4 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SOLEIL SOLEIL  / Beamline: PROXIMA 1 / Wavelength: 0.979 / Beamline: PROXIMA 1 / Wavelength: 0.979 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Apr 19, 2009 / Details: KIRKPATRICK-BAEZ PAIR OF BI-MORPH MIRRORS |
| Radiation | Monochromator: CHANNEL CUT MONOCHROMATOR CRYSTAL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.979 Å / Relative weight: 1 : 0.979 Å / Relative weight: 1 |
| Reflection | Resolution: 1.5→31.7 Å / Num. obs: 28098 / % possible obs: 97.9 % / Observed criterion σ(I): 3 / Redundancy: 6.1 % / Biso Wilson estimate: 17.57 Å2 / Rmerge(I) obs: 0.06 / Net I/σ(I): 16.3 |
| Reflection shell | Resolution: 1.5→1.58 Å / Redundancy: 6 % / Rmerge(I) obs: 0.44 / Mean I/σ(I) obs: 3.6 / % possible all: 96.3 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  SAD SADStarting model: NONE Resolution: 1.5→18.37 Å / Cor.coef. Fo:Fc: 0.9576 / Cor.coef. Fo:Fc free: 0.9524 / SU R Cruickshank DPI: 0.063 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.067 / SU Rfree Blow DPI: 0.065 / SU Rfree Cruickshank DPI: 0.062 Details: IDEAL-DIST CONTACT TERM CONTACT SETUP. RESIDUE TYPES WITHOUT CCP4 ATOM TYPE IN LIBRARY=CL. NUMBER OF ATOMS WITH PROPER CCP4 ATOM TYPE=1314. NUMBER WITH APPROX DEFAULT CCP4 ATOM TYPE=0. ...Details: IDEAL-DIST CONTACT TERM CONTACT SETUP. RESIDUE TYPES WITHOUT CCP4 ATOM TYPE IN LIBRARY=CL. NUMBER OF ATOMS WITH PROPER CCP4 ATOM TYPE=1314. NUMBER WITH APPROX DEFAULT CCP4 ATOM TYPE=0. NUMBER TREATED BY BAD NON-BONDED CONTACTS=1.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 22.93 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.182 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.5→18.37 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.5→1.56 Å / Total num. of bins used: 14
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller





























 PDBj
PDBj




