[English] 日本語
 Yorodumi
Yorodumi- PDB-2pzd: Crystal Structure of the HtrA2/Omi PDZ Domain Bound to a Phage-De... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2pzd | ||||||
|---|---|---|---|---|---|---|---|
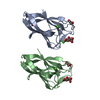
| Title | Crystal Structure of the HtrA2/Omi PDZ Domain Bound to a Phage-Derived Ligand (WTMFWV) | ||||||
 Components Components | Serine protease HTRA2 | ||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  PDZ domain / PDZ domain /  serine protease / serine protease /  apoptosis / apoptosis /  mitochondria / peptide-binding module mitochondria / peptide-binding module | ||||||
| Function / homology |  Function and homology information Function and homology information HtrA2 peptidase / pentacyclic triterpenoid metabolic process / negative regulation of mitophagy in response to mitochondrial depolarization / positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / regulation of autophagy of mitochondrion / ceramide metabolic process / CD40 receptor complex / : / HtrA2 peptidase / pentacyclic triterpenoid metabolic process / negative regulation of mitophagy in response to mitochondrial depolarization / positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / regulation of autophagy of mitochondrion / ceramide metabolic process / CD40 receptor complex / : /  programmed cell death / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand ... programmed cell death / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand ... HtrA2 peptidase / pentacyclic triterpenoid metabolic process / negative regulation of mitophagy in response to mitochondrial depolarization / positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / regulation of autophagy of mitochondrion / ceramide metabolic process / CD40 receptor complex / : / HtrA2 peptidase / pentacyclic triterpenoid metabolic process / negative regulation of mitophagy in response to mitochondrial depolarization / positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / regulation of autophagy of mitochondrion / ceramide metabolic process / CD40 receptor complex / : /  programmed cell death / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand / serine-type endopeptidase complex / adult walking behavior / response to herbicide / positive regulation of protein targeting to mitochondrion / execution phase of apoptosis / positive regulation of cysteine-type endopeptidase activity involved in apoptotic process / protein autoprocessing / regulation of multicellular organism growth / negative regulation of cell cycle / neuron development / cellular response to interferon-beta / negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / forebrain development / cellular response to retinoic acid / serine-type peptidase activity / mitochondrion organization / programmed cell death / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand / serine-type endopeptidase complex / adult walking behavior / response to herbicide / positive regulation of protein targeting to mitochondrion / execution phase of apoptosis / positive regulation of cysteine-type endopeptidase activity involved in apoptotic process / protein autoprocessing / regulation of multicellular organism growth / negative regulation of cell cycle / neuron development / cellular response to interferon-beta / negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / forebrain development / cellular response to retinoic acid / serine-type peptidase activity / mitochondrion organization /  mitochondrial membrane / protein catabolic process / mitochondrial membrane / protein catabolic process /  mitochondrial intermembrane space / cytoplasmic side of plasma membrane / cellular response to growth factor stimulus / intrinsic apoptotic signaling pathway in response to DNA damage / unfolded protein binding / cellular response to oxidative stress / cellular response to heat / mitochondrial intermembrane space / cytoplasmic side of plasma membrane / cellular response to growth factor stimulus / intrinsic apoptotic signaling pathway in response to DNA damage / unfolded protein binding / cellular response to oxidative stress / cellular response to heat /  peptidase activity / neuron apoptotic process / negative regulation of neuron apoptotic process / peptidase activity / neuron apoptotic process / negative regulation of neuron apoptotic process /  cytoskeleton / positive regulation of apoptotic process / serine-type endopeptidase activity / cytoskeleton / positive regulation of apoptotic process / serine-type endopeptidase activity /  chromatin / endoplasmic reticulum membrane / chromatin / endoplasmic reticulum membrane /  endoplasmic reticulum / endoplasmic reticulum /  mitochondrion / mitochondrion /  proteolysis / proteolysis /  membrane / identical protein binding / membrane / identical protein binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.75 Å MOLECULAR REPLACEMENT / Resolution: 2.75 Å | ||||||
 Authors Authors | Appleton, B.A. / Wiesmann, C. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2007 Journal: Protein Sci. / Year: 2007Title: Structural and functional analysis of the ligand specificity of the HtrA2/Omi PDZ domain. Authors: Zhang, Y. / Appleton, B.A. / Wu, P. / Wiesmann, C. / Sidhu, S.S. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE The peptide ligand was fused to the C-terminus of the linker |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2pzd.cif.gz 2pzd.cif.gz | 55.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2pzd.ent.gz pdb2pzd.ent.gz | 40.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2pzd.json.gz 2pzd.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pz/2pzd https://data.pdbj.org/pub/pdb/validation_reports/pz/2pzd ftp://data.pdbj.org/pub/pdb/validation_reports/pz/2pzd ftp://data.pdbj.org/pub/pdb/validation_reports/pz/2pzd | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1lcyS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Unit cell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments: Ens-ID: 1 / Refine code: 3
|
- Components
Components
| #1: Protein | Mass: 12593.536 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: HTRA2 / Production host: Homo sapiens (human) / Gene: HTRA2 / Production host:   Escherichia coli (E. coli) / References: UniProt: O43464, Escherichia coli (E. coli) / References: UniProt: O43464,  HtrA2 peptidase HtrA2 peptidase#2: Chemical |  Ethylene glycol Ethylene glycol#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.36 Å3/Da / Density % sol: 63.43 % |
|---|---|
Crystal grow | Temperature: 292 K / Method: vapor diffusion / pH: 5.6 Details: 0.1 M sodium citrate, 1.0 M monoammonium dihydrogen phosphate, pH 5.6, VAPOR DIFFUSION, temperature 292K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL9-1 / Wavelength: 0.975 Å / Beamline: BL9-1 / Wavelength: 0.975 Å |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.975 Å / Relative weight: 1 : 0.975 Å / Relative weight: 1 |
| Reflection | Resolution: 2.75→50 Å / Num. obs: 9406 / % possible obs: 99.8 % / Redundancy: 6.9 % / Rsym value: 0.071 / Χ2: 1.077 / Net I/σ(I): 11.2 |
| Reflection shell | Resolution: 2.75→2.85 Å / Redundancy: 7.2 % / Mean I/σ(I) obs: 3.6 / Num. unique all: 901 / Rsym value: 0.594 / Χ2: 1.036 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1LCY Resolution: 2.75→20 Å / Cor.coef. Fo:Fc: 0.938 / Cor.coef. Fo:Fc free: 0.902 / SU B: 11.77 / SU ML: 0.228 / TLS residual ADP flag: LIKELY RESIDUAL / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.521 / ESU R Free: 0.31 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 35.281 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.75→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | Dom-ID: 1 / Auth asym-ID: A / Ens-ID: 1 / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.75→2.807 Å / Total num. of bins used: 25
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj


