+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2juj | ||||||
|---|---|---|---|---|---|---|---|
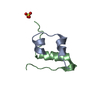
| Title | Solution Structure of the UBA domain from c-Cbl | ||||||
 Components Components | E3 ubiquitin-protein ligase CBL | ||||||
 Keywords Keywords |  LIGASE / LIGASE /  alpha helix / alpha helix /  UBA domain / UBA domain /  Calcium / Calcium /  Cytoplasm / Metal-binding / Cytoplasm / Metal-binding /  Phosphorylation / Phosphorylation /  Proto-oncogene / Proto-oncogene /  SH2 domain / Ubl conjugation pathway / SH2 domain / Ubl conjugation pathway /  Zinc / Zinc /  Zinc-finger Zinc-finger | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of platelet-derived growth factor receptor-alpha signaling pathway / ubiquitin-dependent endocytosis / regulation of Rap protein signal transduction / entry of bacterium into host cell /  flotillin complex / phosphatidylinositol 3-kinase regulatory subunit binding / positive regulation of epidermal growth factor receptor signaling pathway / Regulation of KIT signaling / mast cell degranulation / Interleukin-6 signaling ...regulation of platelet-derived growth factor receptor-alpha signaling pathway / ubiquitin-dependent endocytosis / regulation of Rap protein signal transduction / entry of bacterium into host cell / flotillin complex / phosphatidylinositol 3-kinase regulatory subunit binding / positive regulation of epidermal growth factor receptor signaling pathway / Regulation of KIT signaling / mast cell degranulation / Interleukin-6 signaling ...regulation of platelet-derived growth factor receptor-alpha signaling pathway / ubiquitin-dependent endocytosis / regulation of Rap protein signal transduction / entry of bacterium into host cell /  flotillin complex / phosphatidylinositol 3-kinase regulatory subunit binding / positive regulation of epidermal growth factor receptor signaling pathway / Regulation of KIT signaling / mast cell degranulation / Interleukin-6 signaling / response to testosterone / negative regulation of epidermal growth factor receptor signaling pathway / cellular response to platelet-derived growth factor stimulus / response to starvation / protein monoubiquitination / TGF-beta receptor signaling activates SMADs / protein autoubiquitination / FLT3 signaling by CBL mutants / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / Negative regulation of FLT3 / phosphotyrosine residue binding / flotillin complex / phosphatidylinositol 3-kinase regulatory subunit binding / positive regulation of epidermal growth factor receptor signaling pathway / Regulation of KIT signaling / mast cell degranulation / Interleukin-6 signaling / response to testosterone / negative regulation of epidermal growth factor receptor signaling pathway / cellular response to platelet-derived growth factor stimulus / response to starvation / protein monoubiquitination / TGF-beta receptor signaling activates SMADs / protein autoubiquitination / FLT3 signaling by CBL mutants / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / Negative regulation of FLT3 / phosphotyrosine residue binding /  ephrin receptor binding / InlB-mediated entry of Listeria monocytogenes into host cell / cellular response to nerve growth factor stimulus / response to activity / Regulation of signaling by CBL / Negative regulation of FGFR3 signaling / response to gamma radiation / Negative regulation of FGFR2 signaling / EGFR downregulation / Negative regulation of FGFR4 signaling / Negative regulation of FGFR1 signaling / Spry regulation of FGF signaling / Constitutive Signaling by EGFRvIII / RING-type E3 ubiquitin transferase / Negative regulation of MET activity / ephrin receptor binding / InlB-mediated entry of Listeria monocytogenes into host cell / cellular response to nerve growth factor stimulus / response to activity / Regulation of signaling by CBL / Negative regulation of FGFR3 signaling / response to gamma radiation / Negative regulation of FGFR2 signaling / EGFR downregulation / Negative regulation of FGFR4 signaling / Negative regulation of FGFR1 signaling / Spry regulation of FGF signaling / Constitutive Signaling by EGFRvIII / RING-type E3 ubiquitin transferase / Negative regulation of MET activity /  cilium / cilium /  receptor tyrosine kinase binding / receptor tyrosine kinase binding /  SH3 domain binding / protein polyubiquitination / cytokine-mediated signaling pathway / positive regulation of receptor-mediated endocytosis / ubiquitin-protein transferase activity / male gonad development / Signaling by CSF1 (M-CSF) in myeloid cells / SH3 domain binding / protein polyubiquitination / cytokine-mediated signaling pathway / positive regulation of receptor-mediated endocytosis / ubiquitin-protein transferase activity / male gonad development / Signaling by CSF1 (M-CSF) in myeloid cells /  ubiquitin protein ligase activity / Cargo recognition for clathrin-mediated endocytosis / Constitutive Signaling by Ligand-Responsive EGFR Cancer Variants / ubiquitin protein ligase activity / Cargo recognition for clathrin-mediated endocytosis / Constitutive Signaling by Ligand-Responsive EGFR Cancer Variants /  Clathrin-mediated endocytosis / Clathrin-mediated endocytosis /  growth cone / ubiquitin-dependent protein catabolic process / cellular response to hypoxia / response to ethanol / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / protein ubiquitination / growth cone / ubiquitin-dependent protein catabolic process / cellular response to hypoxia / response to ethanol / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / protein ubiquitination /  cadherin binding / cadherin binding /  membrane raft / membrane raft /  focal adhesion / DNA damage response / focal adhesion / DNA damage response /  calcium ion binding / negative regulation of apoptotic process / perinuclear region of cytoplasm / calcium ion binding / negative regulation of apoptotic process / perinuclear region of cytoplasm /  Golgi apparatus / Golgi apparatus /  signal transduction / signal transduction /  plasma membrane / plasma membrane /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Zhou, Z.R. / Hong, J. / Lin, D.H. / Hu, H.Y. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2008 Journal: Protein Sci. / Year: 2008Title: Differential Ubiquitin Binding of the UBA Domains from Human c-Cbl and Cbl-b: NMR Structural and Biochemical Insights. Authors: Zhou, Z.R. / Gao, H.C. / Zhou, C.J. / Chang, Y.G. / Hong, J. / Song, A.X. / Lin, D.H. / Hu, H.Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2juj.cif.gz 2juj.cif.gz | 253.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2juj.ent.gz pdb2juj.ent.gz | 210.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2juj.json.gz 2juj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ju/2juj https://data.pdbj.org/pub/pdb/validation_reports/ju/2juj ftp://data.pdbj.org/pub/pdb/validation_reports/ju/2juj ftp://data.pdbj.org/pub/pdb/validation_reports/ju/2juj | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 6112.852 Da / Num. of mol.: 1 / Fragment: UBA domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: CBL, CBL2, RNF55 / Production host: Homo sapiens (human) / Gene: CBL, CBL2, RNF55 / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3) Escherichia coli (E. coli) / Strain (production host): BL21(DE3)References: UniProt: P22681,  Ligases; Forming carbon-nitrogen bonds; Acid-amino-acid ligases (peptide synthases) Ligases; Forming carbon-nitrogen bonds; Acid-amino-acid ligases (peptide synthases) |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 50 / pH: 6.5 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Varian Unity / Manufacturer: Varian / Model : UNITY / Field strength: 600 MHz : UNITY / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software | Name: CNS / Classification: refinement |
|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: The structures were calculated with ARIA/CNS using torsion angle dynamics |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: 15 structures for lowest energy Conformers calculated total number: 200 / Conformers submitted total number: 15 |
 Movie
Movie Controller
Controller











 PDBj
PDBj







