Entry Database : PDB / ID : 2jg8Title Crystallographic structure of human C1q globular heads complexed to phosphatidyl-serine (Complement C1q subcomponent subunit ...) x 3 Keywords / / / / / / / / / / / / / / Function / homology Biological species Homo sapiens (human)Method / / / Resolution : 2.05 Å Authors Paidassi, H. / Tacnet-Delorme, P. / Garlatti, V. / Darnault, C. / Ghebrehiwet, B. / Gaboriaud, C. / Arlaud, G.J. / Frachet, P. Journal : J.Immunol. / Year : 2008Title : C1Q Binds Phosphatidylserine and Likely Acts as a Multiligand-Bridging Molecule in Apoptotic Cell Recognition.Authors : Paidassi, H. / Tacnet-Delorme, P. / Garlatti, V. / Darnault, C. / Ghebrehiwet, B. / Gaboriaud, C. / Arlaud, G.J. / Frachet, P. History Deposition Feb 9, 2007 Deposition site / Processing site Revision 1.0 Feb 19, 2008 Provider / Type Revision 1.1 May 8, 2011 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Dec 5, 2018 Group Advisory / Data collection ... Advisory / Data collection / Database references / Source and taxonomy / Structure summary Category citation / entity ... citation / entity / entity_name_com / entity_src_nat / pdbx_entity_src_syn / pdbx_unobs_or_zero_occ_atoms / struct_ref / struct_ref_seq Item _citation.journal_id_ASTM / _citation.journal_id_CSD ... _citation.journal_id_ASTM / _citation.journal_id_CSD / _citation.page_last / _citation.pdbx_database_id_DOI / _entity.pdbx_description / _entity.src_method / _struct_ref.pdbx_align_begin / _struct_ref.pdbx_seq_one_letter_code / _struct_ref_seq.db_align_beg / _struct_ref_seq.db_align_end Revision 1.4 Jul 29, 2020 Group Data collection / Derived calculations ... Data collection / Derived calculations / Other / Structure summary Category chem_comp / entity ... chem_comp / entity / pdbx_chem_comp_identifier / pdbx_database_status / pdbx_entity_nonpoly / pdbx_struct_conn_angle / struct_conn / struct_site / struct_site_gen Item _chem_comp.name / _chem_comp.type ... _chem_comp.name / _chem_comp.type / _entity.pdbx_description / _pdbx_database_status.status_code_sf / _pdbx_entity_nonpoly.name / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id Description / Provider / Type Revision 1.5 Dec 13, 2023 Group Advisory / Data collection ... Advisory / Data collection / Database references / Refinement description / Structure summary Category chem_comp / chem_comp_atom ... chem_comp / chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / pdbx_unobs_or_zero_occ_atoms Item / _database_2.pdbx_DOI / _database_2.pdbx_database_accession
Show all Show less Remark 700 SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN ... SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN THE SHEET RECORDS BELOW, TWO SHEETS ARE DEFINED.
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords IMMUNE SYSTEM / POLYMORPHISM /
IMMUNE SYSTEM / POLYMORPHISM /  GLYCOPROTEIN /
GLYCOPROTEIN /  PHAGOCYTOSIS / DISEASE MUTATION / COMPLEMENT PATHWAY / CELL SURFACE MOLECULE /
PHAGOCYTOSIS / DISEASE MUTATION / COMPLEMENT PATHWAY / CELL SURFACE MOLECULE /  PYRROLIDONE CARBOXYLIC ACID /
PYRROLIDONE CARBOXYLIC ACID /  HYDROXYLATION /
HYDROXYLATION /  INNATE IMMUNITY /
INNATE IMMUNITY /  IMMUNE RESPONSE /
IMMUNE RESPONSE /  COLLAGEN /
COLLAGEN /  TOLERANCE / APOPOTOSIS / COMPLEMENT
TOLERANCE / APOPOTOSIS / COMPLEMENT Function and homology information
Function and homology information complement activation / Classical antibody-mediated complement activation ...complement component C1 complex / complement component C1q complex / negative regulation of macrophage differentiation / synapse pruning / negative regulation of granulocyte differentiation / vertebrate eye-specific patterning / complement-mediated synapse pruning / collagen trimer /
complement activation / Classical antibody-mediated complement activation ...complement component C1 complex / complement component C1q complex / negative regulation of macrophage differentiation / synapse pruning / negative regulation of granulocyte differentiation / vertebrate eye-specific patterning / complement-mediated synapse pruning / collagen trimer /  complement activation / Classical antibody-mediated complement activation / neuron remodeling / Initial triggering of complement /
complement activation / Classical antibody-mediated complement activation / neuron remodeling / Initial triggering of complement /  complement activation, classical pathway /
complement activation, classical pathway /  Regulation of Complement cascade / astrocyte activation / synapse organization / microglial cell activation / cell-cell signaling /
Regulation of Complement cascade / astrocyte activation / synapse organization / microglial cell activation / cell-cell signaling /  amyloid-beta binding / postsynapse / collagen-containing extracellular matrix / blood microparticle /
amyloid-beta binding / postsynapse / collagen-containing extracellular matrix / blood microparticle /  immune response /
immune response /  innate immune response /
innate immune response /  synapse /
synapse /  extracellular space / extracellular region
extracellular space / extracellular region
 Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.05 Å
MOLECULAR REPLACEMENT / Resolution: 2.05 Å  Authors
Authors Citation
Citation Journal: J.Immunol. / Year: 2008
Journal: J.Immunol. / Year: 2008 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2jg8.cif.gz
2jg8.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2jg8.ent.gz
pdb2jg8.ent.gz PDB format
PDB format 2jg8.json.gz
2jg8.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/jg/2jg8
https://data.pdbj.org/pub/pdb/validation_reports/jg/2jg8 ftp://data.pdbj.org/pub/pdb/validation_reports/jg/2jg8
ftp://data.pdbj.org/pub/pdb/validation_reports/jg/2jg8

 Links
Links Assembly
Assembly


 Components
Components
 Homo sapiens (human) / References: UniProt: P02745
Homo sapiens (human) / References: UniProt: P02745
 Homo sapiens (human) / References: UniProt: P02746
Homo sapiens (human) / References: UniProt: P02746
 Homo sapiens (human) / References: UniProt: P02747
Homo sapiens (human) / References: UniProt: P02747
 N-Acetylglucosamine
N-Acetylglucosamine
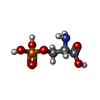



 Phosphoserine
Phosphoserine Water
Water X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  ESRF
ESRF  / Beamline: ID14-2 / Wavelength: 0.933
/ Beamline: ID14-2 / Wavelength: 0.933  : 0.933 Å / Relative weight: 1
: 0.933 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj




