+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qry | ||||||
|---|---|---|---|---|---|---|---|
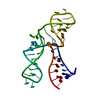
| Title | Homeobox protein VND (ventral nervous system defective protein) | ||||||
 Components Components | PROTEIN (HOMEOBOX VENTRAL NERVOUS SYSTEM DEFECTIVE PROTEIN) | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / DNA BINDING PROTEIN /  HELIX-TURN-HELIX / HELIX-TURN-HELIX /  DNA-BINDING PROTEIN DNA-BINDING PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationneuroblast development / neuroblast fate determination / glial cell development /  brain development / DNA-binding transcription factor binding / brain development / DNA-binding transcription factor binding /  cell differentiation / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / negative regulation of gene expression / regulation of transcription by RNA polymerase II ...neuroblast development / neuroblast fate determination / glial cell development / cell differentiation / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / negative regulation of gene expression / regulation of transcription by RNA polymerase II ...neuroblast development / neuroblast fate determination / glial cell development /  brain development / DNA-binding transcription factor binding / brain development / DNA-binding transcription factor binding /  cell differentiation / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / negative regulation of gene expression / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / cell differentiation / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / negative regulation of gene expression / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Xiang, B. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2003 Journal: Biochemistry / Year: 2003Title: Distortion of the three-dimensional structure of the vnd/NK-2 homeodomain bound to DNA induced by an embryonically lethal A35T point mutation. Authors: Hwang, K.J. / Xiang, B. / Gruschus, J.M. / Nam, K.Y. / No, K.T. / Nirenberg, M. / Ferretti, J.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qry.cif.gz 1qry.cif.gz | 539.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qry.ent.gz pdb1qry.ent.gz | 467.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qry.json.gz 1qry.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qr/1qry https://data.pdbj.org/pub/pdb/validation_reports/qr/1qry ftp://data.pdbj.org/pub/pdb/validation_reports/qr/1qry ftp://data.pdbj.org/pub/pdb/validation_reports/qr/1qry | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9747.162 Da / Num. of mol.: 1 / Fragment: vnd/NK2 homeodomain / Mutation: A35T Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Drosophila melanogaster (fruit fly) / Plasmid: PET-15B / Production host: Drosophila melanogaster (fruit fly) / Plasmid: PET-15B / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE)PLYSS / References: UniProt: P22808 Escherichia coli (E. coli) / Strain (production host): BL21(DE)PLYSS / References: UniProt: P22808 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||
| NMR details | Text: 3D 15N EDITED NOESY-HSQC, 4D 15N/13C EDITED HMQC-NOESY-HSQC, AND 4D 13C/13C EDITED HMQC-NOESY-HSQC WERE USED FOR OBTAINING NOE RESTRAINTS. |
- Sample preparation
Sample preparation
| Details |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | pH: 6.8 / Pressure: AMBIENT / Temperature: 308 K | ||||||||
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker AMX / Manufacturer: Bruker / Model : AMX / Field strength: 600 MHz : AMX / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: THE STRUCTURES ARE BASED ON 1146 NOE-DERIVED DISTANCE RESTRAINTS AND 43 DIHEDRAL ANGLE RESTRAINTS. | ||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||
| NMR ensemble | Conformer selection criteria: STRUCTURES WITH ACCEPTABLE COVALENT GEOMETRY,STRUCTURES WITH THE LEAST RESTRAINT VIOLATIONS,STRUCTURES WITH THE LOWEST ENERGY Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller











 PDBj
PDBj
